What is cascade testing?
Cascade testing is the practice of offering genetic testing to relatives of known carriers of pathogenic variants associated with autosomal dominant conditions. In oncology, cascade testing is performed in families affected by hereditary cancer syndromes1. The most common include hereditary breast and ovarian cancer syndrome (HBOC), Lynch syndrome (LS), familial adenomatous polyposis syndrome, hereditary pancreatic cancer syndrome and gastric cancer syndrome2,3. Testing for variants associated with HBOC and LS belongs to the so-called Tier 1 testing, i.e., genomic applications, the implementation of which is supported by robust evidence4.
Cascade testing starts with first-degree relatives (parents, siblings, children) of index cases (i.e., the family member in whom a pathogenic variant was identified) and then proceeds to second- (grandparents/grandchildren, aunts/uncles, nieces/nephews, half-siblings) and third-degree relatives (great-grandparents/great-grandchildren, first cousins)1.
Most hereditary cancer syndromes follow the autosomal dominant inheritance pattern. Therefore, the first-, second- and third-degree relatives have, respectively, a 50%, 25%, and 12.5% probability of inheriting the predisposition to develop cancer (see Figure 1)1. For some pathogenic variants of genes associated with hereditary cancer syndromes, such as BRCA1/2, the penetrance is high5. Establishing accurate estimates of penetrance and relative risk for genes implicated in hereditary cancer syndromes is an ongoing task3.
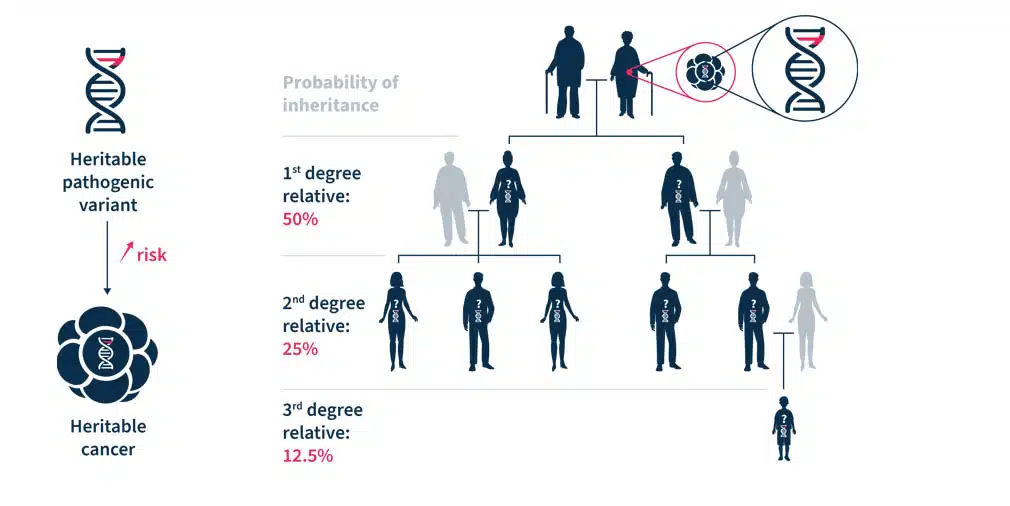
Figure 1. Heritable pathogenic variants increase the risk of developing cancer at a younger age (left). Cascade genetic testing is the practice of testing the relatives of known carriers (right)1.
Why is cascade testing important?
At the level of an individual and their family, cascade testing has two important goals. The first goal is to identify relatives that carry the familial pathogenic variant and require personalized cancer risk management1. The second goal is to exclude the non-carriers from intensive cancer surveillance and prevention interventions1. The detection of pathogenic variants in individuals at a reproductive age may lead to decisions of assisted reproduction or prenatal diagnosis. In the case of actionable monogenic conditions, cascade testing may reduce adverse health outcomes in cohorts of relatives1.
At the societal level, cascade testing has important clinical and research implications for oncology. It can further our knowledge of hereditary cancers and is a cost-effective way of identifying unaffected individuals at-risk, thus, providing important information to plan long-term resources necessary to cope with hereditary cancers. Moreover, today’s testing is needed to tailor future approaches in cascade testing1.
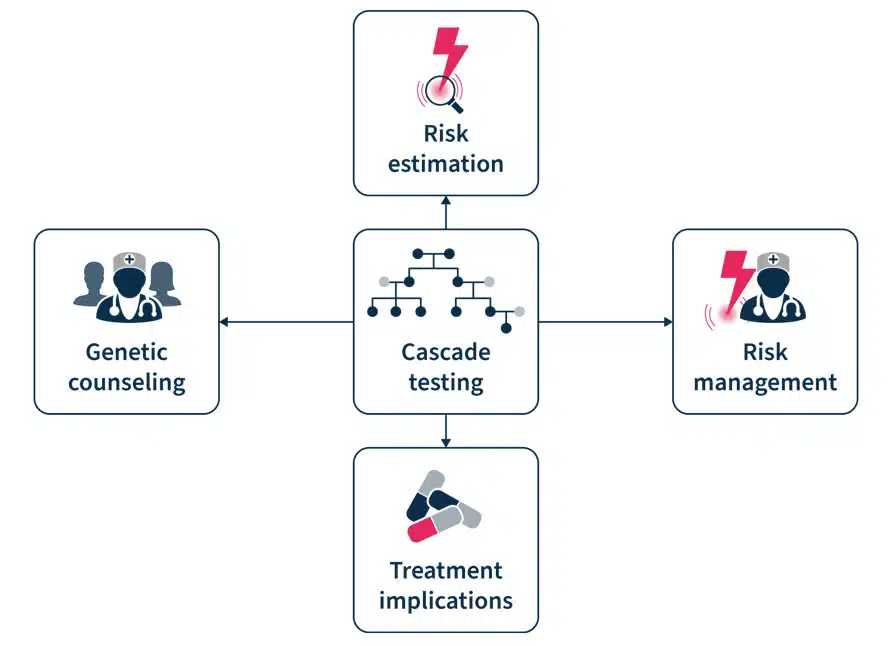
Figure 2. Cascade testing involves genetic counseling before and after the test, risk estimation and management, and has treatment implication7.
What are the barriers to cascade testing?
Despite the advantages of cascade testing, its uptake is low. The reported rates of uptake of cascade testing in HBOC and LS equals ~50% and the underutilization of testing results in missed opportunities of cancer prevention1. In a recent Swiss study, there was a 25-50% response rate to invitations to cascade testing and at least one-in-three individuals at risk did not undergo testing. An index case possesses an average of 10 relatives eligible for testing, while the average rate of genetic tests per index case is only 1.51.
There are several barriers to cascade testing6,7. These include ineffective family communication of genetic risk information, low knowledge of cascade testing among index cases and primary care providers, and geographic barriers to receiving genetic services. Cascade testing uptake is also lower among male than female relatives and in distant compared to first-degree relatives. A facilitator of adherence to cascade testing is the parents’ desire to understand their children’s risk6. “Dear family” letters, digital chatbots (a technology-based simulated conversations), and direct contact programs have been shown to be effective in motivating cascade testing8.
Several initiatives exist to promote cascade testing. One such enterprise is the Cascade Resources Network, an independently run, non-profit organization that offers access to genetic testing, genetic counseling, variant interpretation, screening guidelines, and forums and support. It was developed by Memorial Sloan Kettering Cancer Center (MSK) fellows, Ryan Kahn and Sushmita Gordhandas. The network was created to increase the rate of genetic testing among relatives of patients with inherited cancer risk variants to help identify cancer early in families and, ultimately, to prevent future cancers. Similarly, the Swiss Cancer Genetic Predisposition Cascade Screening Consortium was established in 2016 to foster research related to the hereditary cancer predisposition. In particular, the Consortium promotes the CASCADE cohort, a family-based open-ended cohort targeting HBOC and LS variant-harboring families to elicit factors that enhance adherence to testing (NCT03124212).
Analyze genetic predisposition to cancer with the SOPHiA DDMTM Platform
Multi-gene testing is an efficient, affordable, and guideline-recommended9 approach to cascade testing as it allows for comprehensive assessment of biologically relevant hereditary cancer genes. The SOPHiA DDM™ Platform supports various next generation sequencing (NGS)-based Hereditary Cancer Applications to help clinician researchers characterize the complex mutational landscape associated with hereditary cancer disorders.
Powered by advanced analytics, users can detect challenging variants in a streamlined sample-to-report workflow, including:
- Single nucleotide variants (SNVs), insertions and deletions (Indels) and copy number variations (CNVs)
- Boland inversions
- Alu insertions
- PMS2 and PMS2CL variants
Variant pathogenicity levels are assigned using machine learning complemented by guideline-driven ranking, helping to prioritize relevant variants and reduce interpretation time. Furthermore, deeper variant exploration is supported by Alamut™ Visual Plus, a full-genome browser that integrates numerous curated genomic and literature databases, guidelines, missense and splicing predictors.
To learn more about SOPHiA DDM™ for Hereditary Cancers, explore here or request a demo here.
References
1. Sarki M, et al. Cancers (Basel) 2022;14:1636.
2. Brown GR, et al. JAAPA 2020;33(12):10-16.
3. Mighton C, Lerner-Ellis JP. Genes Chromosomes Cancer 2022;61(6):356-381.
4. Dotson WD, et al. Clin Pharmacol Ther 2014;95(4):394-402.
5. Chen S, Parmigiani G. J Clin Oncol 2007;25(11):1329-1333.
6. Roberts MC, et al. Health Aff (Millwood) 2018;37(5):801-808.
7. O’Neill SC, et al. Hered Cancer Clin Pract 2021;19(1):40.
8. Campbell-Salome G, et al. (2022) Transl Behav Med 2022;12(7):800–809.
9. Daly MB, et al. J Natl Compr Canc Netw. 2021 Jan 6;19(1):77-102.