Menu
Menu



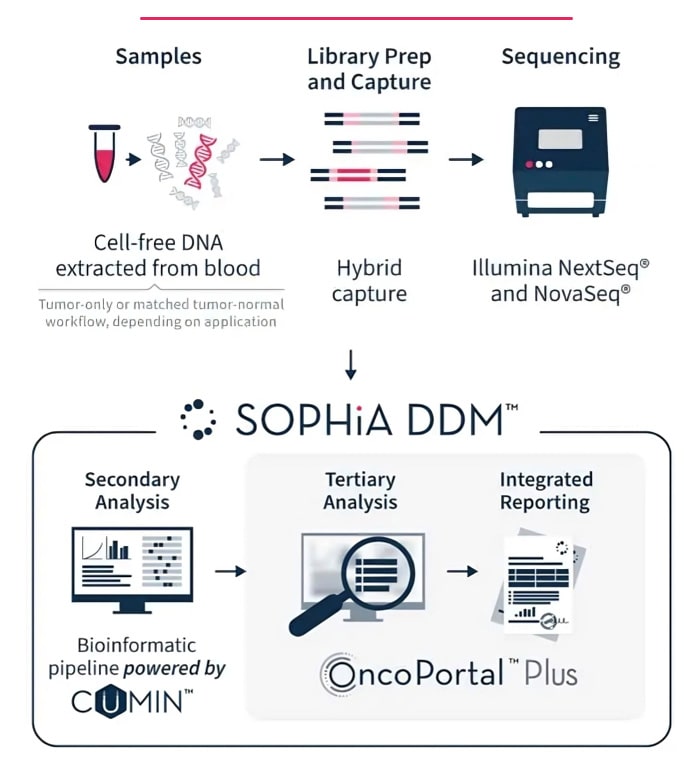
Rely on robust sequencing across multiple applications using one automated protocol.
Access variant ranking, virtual panels, and familial analysis.
Reveal previously overlooked variants with the Alamut™ Visual Plus full genome browser.
Clearly summarize key variants and supporting information in tailored reports.
Sensitive SNV and Indel detection
Accurate CNV calling
Efficient star allele calling
Robust variant annotation
Smart variant prioritization



Achieve streamlined and accurate variant detection and interpretation in a single experiment by combining SOPHiA DDM™’s advanced analytical capabilities with Alamut™ Visual Plus’ enhanced annotation and visualization, with the optimal exome or genome sequencing enrichment kit and sequencer for your lab.
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™ Dx Hereditary Cancer Solution, SOPHiA DDM™ Dx RNAtarget Oncology Solution and SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution are available as CE-IVD products for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA DDM™ Dx Myeloid Solution and SOPHiA DDM™ Dx Solid Tumor Solution are available as CE-IVD products for In Vitro Diagnostic Use in the EEA, the United Kingdom, Switzerland, and Israel. Information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us to obtain the appropriate product information for your country of residence.
All third-party trademarks listed by SOPHiA GENETICS remain the property of their respective owners. Unless specifically identified as such, SOPHiA GENETICS’ use of third-party trademarks does not indicate any relationship, sponsorship, or endorsement between SOPHiA GENETICS and the owners of these trademarks. Any references by SOPHiA GENETICS to third-party trademarks is to identify the corresponding third-party goods and/or services and shall be considered nominative fair use under the trademark law.