Menu
Menu
At ASCO 2022, oncology professionals gathered in Chicago, Illinois, and online to discuss the latest advances in research and care for patients with cancer. This year's program featured over 200 sessions on Advancing Equitable Cancer Care Through Innovation. The presentations spanned from care delivery and regulatory policy to developmental therapeutics, gastrointestinal cancer, lung cancer, pediatric oncology, and beyond. Here, discover our summary of four outstanding ASCO presentations focusing on breast cancer treatment, diagnosis and follow-up, showcasing the power of precision medicine in healthcare.
Targetable genomic mutations in young women with advanced breast cancer1.
Norin Ansari, Yale New Haven Hospital, New Haven, CT
Advanced breast cancers (BC) in young women (under 40 years old) are often more aggressive and with worse prognoses than in older women. As treatment strategies can be dictated by the type of genomic alteration (GA), knowledge of BC genetic profiles across ages can greatly improve guidance and outcomes. In her poster presentation, Norin Ansari analyzed over 2,000 BC using hybrid-capture based comprehensive genomic profiling (CGP) to evaluate subtypes of GA and confirmed via immunohistochemistry (IHC) hormone receptors (HR) and PD-L1 status.
The study showed a mutations stratification within the population of BC depending on patient's age. Indeed, younger patients had higher rates of BRCA1, BRCA2, and RB1 mutations and lower rates of CDH1 and PIK3CA mutations than did older patients. Differences were statistically significant in BRCA1, CDH1, and PIK3CA. Norin Ansari also showed that breast tumors in younger women were less likely to be estrogen receptor positive (ER+) and more likely to be triple negative while no clear age-related pattern for HER2 status could be highlighted. Finally, younger women were more often PD-L1 positive and had lower tumor mutational burden (TMB) than their older counterparts.
Different mutational profiles may support differential use of targeted and immune therapies.
With increasing availability of targeted and immune therapies, knowing which GA each group of women has allows to better tailor therapies and leads to more effective treatments. For instance, BRCA1 mutations may lend to PARP inhibitor use while PIK3CA mutations may indicate the benefit of alpelisib prescription. The difference in genetic mutations between age groups can give a head start when treating women with breast cancer and CGP can refine the approach for better results.
Alpelisib + fulvestrant in patients with hormone receptor–positive, human epidermal growth factor receptor 2–negative advanced breast cancer: Biomarker analyses by next-generation sequencing from the SOLAR-1 study2.
Dejan Juric, Massachusetts General Hospital Cancer Center, Harvard Medical School, Boston, MA
PIK3CA mutations account for approximately 40% of the hormone receptor positive (HR+), HER2-negative (HER2-) advanced BC. PIK3CA encodes for a subunit of PI3K, key of a highly interconnected pathway regulating growth and cell survival. PI3K pathway alterations are associated with endocrine therapy resistance, hence the poor prognosis for HR+, HER2– advanced BC.
Dejan Juric introduced the SOLAR-1 phase 3 study, a randomized controlled study testing the efficacy of the combined administration of alpelisib (ALP, a PI3Kα-selective inhibitor and degrader) and fulvestrant (FUL, a selective estrogen receptor degrader) in HR+, HER2- advanced BC patients. SOLAR-1 shows improved progression-free survival (PFS) in ALP + FUL treated patients versus placebo + FUL of 11.0 and 5.7 months, respectively. Going one step further, they measured the efficacy outcomes in patients with specific gene alterations (GA) in a PIK3CA-altered cohort, applying a retrospective exploratory biomarker analysis.
SOLAR-1 baseline tumor samples were tested by next-generation sequencing (NGS) and clinical benefit was assessed using PFS and hazard ratio based on TMB and GA status in the PIK3CA-altered cohort. While ALP + FUL clinical benefit was seen across TMB quartiles, it was more pronounced in patients with a low TMB (PFS of 18.5 months with ALP versus 3.2 months with placebo). They also observed that, despite improved PFS with ALP + FUL treatment in all PIK3CA-altered patients, the level of benefit may depend on the mutation status of other genes involved in MAPK pathway, PI3K pathway, in endocrine therapy or CDK4/6 inhibitors resistance. For instance, greater benefit was observed with altered FGFR1/2 but limited in MYC- and RAD21-altered cohorts. Besides, ALP + FUL efficacy was independent of GA in TP53, ESR1, CCND1, MAP3K1 and ARID1A.
Clinical benefit of ALP + FUL was maintained regardless of alterations in most biomarkers.
To conclude, Dejan Juric showed that ALP + FUL treatment was beneficial in patients with HR+, HER2– advanced BC, especially with a low TMB, but that a comprehensive understanding of the unique mutational profile of each tumor via biomarkers analysis may explain the level of success and thus dictate further care.
Trastuzumab deruxtecan versus treatment of physician’s choice in patients with HER2-low unresectable and/or metastatic breast cancer: Results of DESTINY-Breast04, a randomized, phase 3 study3.
Shanu Modi, Memorial Sloan Kettering Cancer Center, Memorial Hospital, New York, NY, USA
Metastatic breast cancers (mBC) are classified according to the detection of certain receptors in the tumor cells, dictating the type of treatment to offer to the patients. Thus, mBC with an abnormally high quantity of human epidermal growth factor receptor-2 (HER2+) benefit from therapies targeting HER2 protein with monoclonal antibodies, while HER2- mBC receive treatment based on their HR status. However, the dichotomy between HER2+ and HER2- mBC does not suffice to find effective therapies for patients with low level of HER2 (HER2-low) currently treated as HER2-. The limited options and modest benefits of chemotherapy confirm the need for an adapted targeted strategy.
Trastuzumab deruxtecan (T-DXd) is part of a new generation of antibody-drug conjugates that delivers precision-focused chemotherapy directly to the cancer cells. Its activity was shown in tumors across a broad range of HER2 expression and a phase 1 trial showed promising efficacy of T-DXd in patients previously heavily treated with HER2-low mBC. Here, Shanu Modi presented us the DESTINY-Breast04 study, the first randomized phase 3 study of T-DXd for HER2-low mBC and its auspicious results.
Measuring the median progression-free survival (mPFS) in HR-positive mBC patients as primary endpoint, they observed statistically significant and clinically meaningful improvement for patients with HER2-low mBC compared to standard chemotherapy (mPFS 10.1 versus 5.4 months respectively: p<0.0001). Similar benefit was seen in all patients, regardless of their HR status, with T-DXd treatment through PFS and overall survival (OS) compared to standard chemotherapy.
DESTINY-Breast04 establishes HER2-low mBC as a targetable patient population with T-DXd as a new standard of care, with the potential to improve the survival for ~50% of all mBC patients.
We expanded the benefits of HER2 targeted therapy to a new population of breast cancer patients and established T-DXd as the new standard of care for HER2-low mBC.
These ground-breaking results presented at the 2022 ASCO Annual Meeting, and simultaneously published in the New England Journal of Medicine4, were acknowledged with a standing ovation from the audience of specialists. Anticipating these results to be practice changing, the study gives hope for many oncology professionals and patients.
Circulating tumor DNA and late recurrence in high-risk, hormone receptor–positive, HER2-negative breast cancer (CHiRP)5.
Marla Lipsyc-Sharf, Dana-Farber Cancer Institute, Boston, MA
Over half of metastatic recurrences in HR+ BC are late (occurring over 5 years from diagnosis) and thought to arise from minimal residual disease (MRD), a small number of cancer cells left in the body after treatment, hence the benefit of adjuvant therapy. MRD detection via circulating tumor DNA (ctDNA) is associated with high risk of BC recurrence in the early adjuvant setting across tumor subtypes. Little is known, however, about ctDNA for later settings.
Marla Lipsyc-Sharf presented the CHiRP prospective study of late recurrence in patients with high-risk HR+ BC without prior recurrence. 83 patients were followed with whole exome sequencing on primary tumor samples and plasma collection every 6-12 months to be processed with personalized RaDaRTM assay (12-51 variants) to detect ctDNA. Patients did not undergo routine surveillance body imaging or other circulating biomarkers testing. 68.7% of the patients had stage 3 disease and most received chemotherapy (90.4%) and adjuvant endocrine therapy (100%).
8 of 83 (10%) patients had detectable ctDNA at any timepoint during this study. As of last follow-up, 6 of them developed metastatic recurrence at various sites, 6-14 years after primary diagnosis, and one patient with detected ctDNA developed a locoregional recurrence.
All distant recurrences were detectable via ctDNA prior the recurrence with a median lead time of ~1 year.
Despite the low yet steady rate of recurrence in this small cohort with limited follow-up and infrequent plasma sampling (every 6-12 months), this study, published in Journal of Clinical Oncology6, shows that liquid biopsy can provide precious indication on the risk of relapse and thus point towards earlier intervention after MDR detection, improving patients' survival and quality of life.
1 https://meetings.asco.org/abstracts-presentations/210258
2 https://meetings.asco.org/abstracts-presentations/209230
3 https://meetings.asco.org/abstracts-presentations/209021
4 Modi S, Jacot W, Yamashita T, et al. Trastuzumab Deruxtecan in Previously Treated HER2-Low Advanced Breast Cancer [published online ahead of print, 2022 Jun 5]. N Engl J Med. 2022;10.1056/NEJMoa2203690. doi:10.1056/NEJMoa2203690
5 https://meetings.asco.org/abstracts-presentations/209216
6 Lipsyc-Sharf M, de Bruin EC, Santos K, et al. Circulating Tumor DNA and Late Recurrence in High-Risk Hormone Receptor-Positive, Human Epidermal Growth Factor Receptor 2-Negative Breast Cancer [published online ahead of print, 2022 Jun 4]. J Clin Oncol. 2022;JCO2200908. doi:10.1200/JCO.22.00908
Several RNA alterations have been described in Oncology, including gene fusions, recognized driver mutations in neoplasia1. More than 10,000 gene fusions have already been identified in human cancers1; it is estimated that up to 80% of solid tumors could benefit from gene fusion testing2. The number of new drugs specifically targeting gene fusion-positive cancers keeps growing: the advantages that proper gene fusion detection could bring to clinical cancer management are noticeable.
Transcriptome sequencing has emerged as an effective method to identify gene fusions and has become a routine task in cancer research and precision medicine3. However, although a variety of computational tools have been developed over the years, an optimal solution with high analytical performance for fusion detection and the ability to maximize the insights from small precious RNA samples has been lacking.
SOPHiA DDM™ RNAtarget Technology addresses these requirements by combining powerful novel (partner-agnostic) fusion detection capabilities as well as SNV/Indels detection in selected genes and expression changes assessment. Powered by a deep learning algorithm, the Technology works with a very low sample input, a fully customizable gene panel, and a streamlined automated workflow that supports all stages of the analysis with high sensitivity. Finally, a convenient yet powerful and fast results visualization and interpretation are ensured by the associated SOPHiA DDM™ Platform.
To better understand how SOPHiA GENETICS developed the SOPHiA DDM™ RNAtarget Technology and its features, we sat down with Mikhail Pertziger, the Clinical Application Product Manager for SOPHiA DDM™ RNAtarget Technology at SOPHiA GENETICS.
After studying Biomedical Science for my undergraduate degree, I continued with a PhD in the Molecular Biology of Breast and Colon Cancers, so Cancer and Molecular Diagnostics is very much the area where I have spent a lot of my academic and industry years. They say that the 21st century is the era of biology, and I will have to add that precision medicine is the future of cancer management. Biomarker-guided therapies are introducing dramatic differences in how oncology conditions are managed. Fusions are the latest frontier to receive broad applicability in the clinic with more and more drugs being introduced – this is bound to grow and accelerate. I'm excited to be working on a product that allows our customers to have access to a technology that takes on the learnings of the previous years and enables them to be more confident about detecting fusions.
I have worked at SOPHIA GENETICS for just over three years and currently lead the SOPHiA DDM™ RNAtarget Technology development and launch. The development team includes experts in a broad range of applications, including the core development of BioInformatics, NGS, programming, as well as logistics, Regulatory, Legal, and Marketing.
As I mentioned previously, gene fusions are the latest type of biomarker to receive broad applicability in cancer management. The results of targeting fusions reported in clinical trials and now being seen in routine care are fascinating. The number of new drug approvals in fusion-positive cancers has been continuously increasing over the last decade – up to 80% of solid tumors NGS tests could benefit from the inclusion of fusion testing. With histology-agnostic approvals, this number approaches 100%2. In parallel, more clinical trials are being rolled out to target fusion-positive cancers, hopefully leading to further improvements in treatment options in the near future.
Up to 80% of solid tumors NGS tests could benefit from the inclusion of fusion testing2
SOPHiA DDM™ RNAtarget Technology came about from our users' feedback on the need to have an application that allows them to detect novel fusions without sacrificing sensitivity in smaller biopsy samples. The work on this Technology started more than a year ago and has involved many feasibility and optimization studies to ensure that we're not just providing a regular solution, but a product that really helps users achieve more.
What we wanted to provide with this product was the ability for users to have a high-performance fusion detection that could be run in a very small amount of material, a streamlined (but robust) workflow, as well as the ability to not only detect fusions but also be able to extract as much information as possible from that small sample with detection of SNVs and Expression changes. For convenience, the gene content can be customized to fit the lab's individual needs, automated workflow to reduce resource constraints, and, finally, the product runs on the industry-leading SOPHiA DDM™ Platform, providing convenient visualization, annotation, and reporting of the results.
SNV detection in RNA is an intriguing area of development. There are several applications where SNV detection is beneficial, including the ability to run an RNA-only workflow in cases where genes of interest are sufficiently highly expressed. SNV detection in RNA also opens up the possibility of running RNA and DNA workflow sequentially, where the initial RNA workflow will likely detect the majority of relevant variants, leaving only a subset of samples that need to be also processed through the DNA workflow.
Another benefit of detecting SNVs in RNA is the ability to use them as an additional data point in calling SNVs in DNA or using RNA-based SNVs as a backup in case of issues with DNA.
Finally, having the information on SNV VF% in RNA, is like adding an additional dimension to the molecular profile created by DNA SNVs – as this provides more dynamic, rich, and potentially more insightful information on the state of things in the tumor.
Overall, there are many novel and unique ways this feature of SOPHiA DDM™ RNAtarget Technology could be used, and I'm excited to see how our users will utilize it.
One of the primary applications for the product is, of course, Lung cancer because of the limited amount of biopsy material that is generally available and a high number of clinically relevant fusions in this pathology. At the same time, the solution can be deployed to test any solid tumor, and we're looking at the possibility of running it in blood tumors as well. Moreover, because the gene content is entirely customizable, users can tailor the gene content to the needs of their labs, clinical research, or clinical trials that they want to be part of. This makes the application very versatile while removing the obstacle of manually optimizing the pipeline's performance because the SOPHiA GENETICS BioIT team will take care of that. Given the proliferation of tumor-agnostic biomarkers and, in particular, tumor agnostic fusions, the applicability of this Technology is only going to grow and expand in the future, and the fusion detection would become an integral part of genomic profiling of any cancer, together with SNV and CNVs.
SOPHiA DDM™ RNAtarget Technology can be deployed to test any solid tumor
It's really the combination of 3 main features that make for a cohesive and well-rounded product that offers a lot of value from several perspectives - Novel fusion detection, High sensitivity at low input amounts, and Customizability of the panel. These features provide an excellent foundation for a future-proof, high-performance solution. If you look at the market, there are very functional solutions that offer one or two of these features, but not all 3.
In addition to those three features, we included other functionality that I refer to as "two data points for every variant type," where 5’-3' imbalance serves as an additional data point for calling fusions, SNV detection in RNA can be used together with SNV calls in DNA, and expression changes provide further details and reassurance for the CNVs.
Finally, the streamlined protocol that can also be automated further refines the convenience factor of this solution.
"How" is straightforward: SOPHiA DDM™ RNAtarget Technology utilizes a hybrid-capture approach, which targets the key clinically relevant kinases. This protocol is augmented by a careful probe design process to make sure the panel performs at the highest level in the hands of our users.
On the other hand, it is worth highlighting that the detection of novel (or partner-agnostic) fusions is becoming a more and more prominent feature requested by labs striving to provide the highest level of care. This is underpinned by the higher inherent sensitivity, which is becoming more important in the rise in approval of fusion targeting therapies in a partner agnostic manner.
I would say it comes down to the challenges of needing to have high sensitivity in low sample input, the ability to detect novel fusions, and having a tailored solution that perfectly fits the needs of the lab, plus addressing the need for a convenient yet powerful visualization and interpretation platform – the SOPHiA DDM™ Platform.
Microsatellites (MS) are repetitive nucleotide sequences distributed at millions of loci across our genome1. Because of their repetitive sequence, they are prone to DNA polymerase slippage events during DNA replication, resulting in repeat length variations1. Proteins in the highly conserved mismatch repair (MMR) pathway usually identify and repair these types of errors, safeguarding the genome from potentially deleterious mutations2. If this repair machinery fails, for example, due to genetic or epigenetic loss of MMR protein function, the rate of spontaneous mutations on MS regions throughout the genome increases1. Therefore, microsatellite instability (MSI) is considered a molecular fingerprint for defects in the mismatch repair system (dMMR) (Figure 1) and is associated with various malignancies1.

A recent study on more than 11,000 tissue samples covering 39 cancer types showed that microsatellite instability (MSI) was present within 27 cancer types (3.8% of all cancers analyzed). Twelve cancer types were found with an MSI prevalence more significant than 1%, represented mainly by endometrial (31.4%), gastric (19.1%), and colorectal (CRC) adenocarcinomas (16.0%)3. Most MSI tumors arise sporadically4, and other cases result from inherited cancer predisposition syndromes such as Lynch syndrome (LS)5.
MSI has several implications in the management of patients with solid cancers4. First, detection of MSI can be used as a screening strategy for LS, as almost 95% of the LS-associated malignancies have MSI 5. Secondly, MSI status represents a positive prognostic value for localized CRC and gastric cancers6. Finally, in recent years, MSI has gained considerable attention as a predictive marker of immune checkpoint inhibitor therapy response across multiple tumor types5,6.
Identification of MSI relies on comparing the size of selected microsatellites in normal and tumor tissue of the same individual. The extent of the repeat length variation determines the MSI level of the assessed tumor, which can then be classified as MS-instable or MS-stable6.
Despite its recognized pan-cancer value, commonly used methods only support MSI detection in colorectal cancers, since their sensitivity in other tumor types is low. In addition to the tissue-specific differences that impact the sensitivity of MSI detection, the analytical performance is also impacted by patient ethnicity, tumor content, and other sample-specific properties3.
Immunohistochemistry (IHC) and PCR-based assays performed on tumor tissue samples represent the two current reference methods for MSI/dMMR status detection. Being MSI strictly correlated with the proper functioning of the MMR pathway, the immunohistochemistry (IHC) detection of MMR proteins allows to assess MSI and thus determine MS stable or unstable cases1. Low cost and wide availability are the main advantages of this method. However, this test cannot be combined with other molecular diagnostics tests and is also known to provide false positive and negative results due to different technical and biological factors3, 8.
A complementary and rather preferred tool to investigate MSI is polymerase chain reaction (PCR) analysis followed by capillary electrophoresis analysis, which amplifies informative selected MS and then identifies fragment length polymorphisms2. For the PCR to be indicative, standardized panels of highly unstable mononucleotide or dinucleotide repeats should be used. In the case of colorectal cancers, the Bethesda/ National Cancer Institute (NCI) panel of five repeats has been considered the gold standard for many years9. However, because of the extensive tumor specificity of MSI targets, the use of the same small subset of markers becomes inadequate for other cancer types1.
Next-generation sequencing (NGS)-based MSI detection methods allow simultaneous analysis of a larger number of microsatellite regions for each patient with a higher sensitivity for low prevalence mutations2. In addition, NGS enables the analysis of other cancer-related molecular signatures and genetic lesions with one unique assay, facilitating the adoption of MSI clinical testing and supporting personalized cancer therapy8. Despite these advantages, the analysis of MSI through NGS is challenging. NGS accuracy can be low by analyzing tumor-only data due to the inter-and intra-tumor differences in the frequency and position of MSI diagnostic events2. These differences limit the analytical performance of this method, which relies on a baseline reference distribution to determine MSI samples. Several computational approaches have been developed until now to analyze MSI data coming from NGS, considering the difficulties of microsatellite sequences, including management of stutter peak formation induced by PCR amplification during the library preparation step, sequencing errors induced by homopolymers, and the shortcomings of sequencing read length that limit the length of the microsatellites analyzed (Figure 2).
Therefore, NGS-based MS assessment requires expensive and time-consuming bioinformatic analysis with proper algorithms for data interpretation9.
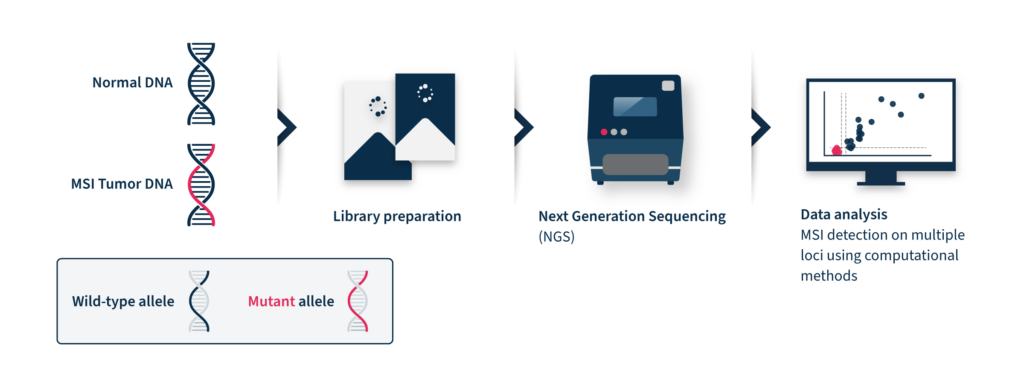
At SOPHiA GENETICS, we believe that data-driven medicine has the potential to improve health outcomes. That’s why we developed MUSTARD™, an algorithm with comparable sensitivity to PCR-based methods and only requiring tumor data to function. MUSTARD™ is a curve-fitting algorithm that aims at better identifying differences in the read length distribution of MSI and normal samples.
MUSTARD™ was effectively developed to perform MSI classification in multiple cancers and trained on a specific selection of more than 100 microsatellite regions identified as highly performant in different cancer types. This means that, across several tumors, the length of these loci significantly differed from MS stable samples. Using such a subset of loci as a reference, the analytical performance of MUSTARD™ increased and became comparable to that of other NGS-based solutions with the great advantage of requiring tumor data only for the analysis.
MUSTARD™ is already available in the SOPHiA DDM™ for TSO500 bioinformatic workflow.
SOPHiA DDM™ has been indeed optimized to detect SNVs, Indels, CNVs, and gene fusions simultaneously to evaluate MSI and tumor mutational burden (TMB), covered by the Illumina TruSight® Oncology 500 (TSO500). The MSI algorithm has been extensively tested on 74 samples, with known MSI status from at least 9 cancer types and 7 different centers. The internal analytical validation study showed 100% sensitivity and specificity, reproducing the MMR/MSI status detected by IHC. Moreover, in a comparison study, MUSTARD™ outperformed the current standard analytical technology for TSO500. In contrast to the standard analytical technology score, which does not allow clear separation between samples classified by IHC as dMMR and pMMR (horizontal line), MUSTARD™ score distribution for these two populations is distinct (vertical lines), (Figure 3).
MUSTARD™ has proven to overcome the main challenges currently faced by most NGS-based solutions and make accurate and extensive MSI detection more attainable than ever. Read more about our MSI detection approach in our Frontiers in Oncology publication.

Inhibition of poly (ADP-ribose) polymerase (PARP) activity induces synthetic lethality in BRCA-mutated tumors by selectively targeting tumor cells that fail to repair DNA double-strand breaks.1 The term ‘BRCAness’ has been coined to describe tumors that are BRCA wild-type but are still sensitive to treatment with PARP inhibitors (PARPi). Like BRCA-mutated tumors, BRCAness tumors have HRD but have mutations in other genes involved in the homologous recombination repair (HRR) pathway (for example, RAD51, CHEK1, CHEK2, BRIP1, ATM). Detecting aberrations in HRR relevant genes, not only BRCA1 and BRCA2, is important to optimize the use of PARPi.1
In a single assay, the SOPHiA DDM™ HRD Solution combines identification of mutations in 28 HRR genes with a measure of genomic integrity called the Genomic Integrity Index. Powered by deep learning algorithms, the Genomic Integrity Index reveals the extent of genomic scarring as a result of HRR gene mutations across the entire genome.
To gain a better understanding of how SOPHiA GENETICS developed the HRD technology, we sat down with Dr. Christian Pozzorini, the Technical Product Manager for the SOPHiA DDM™ HRD Solution and the Director of Biostatistics Research at SOPHiA GENETICS.
After studying life sciences and technology with a focus on genetics and molecular biology during my undergraduate years, I went on to do a PhD in computational neuroscience, a field closely related to machine learning. I have worked at SOPHiA GENETICS for nearly 6 years and currently lead the Biostatistics Research team. The team has extensive expertise in statistical techniques and machine learning. We investigate different methods by which to extract the maximum value out of next generation sequencing (NGS) data, to best support clinical researchers. Two years ago, we recognized HRD and genomic scarring as a key topic in which machine learning technologies could be beneficial to exploit information hidden in NGS data.
HRD is a hot topic in cancer research and is particularly relevant in clinical practice since novel therapies, such as PARPi, have been developed to target HRD tumors. These drugs were initially prescribed to patients with BRCA-deficient tumors, but it is now clear that some BRCA wild-type patients can benefit from the same treatments. The drugs were indeed conceived to target tumors in which the HRR pathway is deficient, and BRCA mutations are only one subset of mutations leading to HRD. HRD testing is particularly relevant in ovarian cancer patients where HRD is frequent, and where HRD testing has been associated with clinical response to PARPi. Additionally, research has shown that other cancer types may have HRD and so HRD testing may also become standard practice when determining treatment options for these cancers.
HRD testing is particularly relevant in ovarian cancer patients where HRD is frequent, and where HRD testing has been associated with clinical response to PARPi.
Detecting HRD is made complicated by the fact that multiple genomic mutations may lead to HRD. If a mutation in a known HRR gene is identified, it is difficult to predict its impact on HRD status. For example, does a particular mutation in PTEN impair the HRR pathway?
To overcome this challenge the scientific community proposed a different, yet complementary, approach. Instead of detecting HRD by looking for genomic aberrations known to cause HRD, rather look for genomic aberrations (or genomic scars) that occur as a result of HRD. The challenge of genomic scarring is that it does not happen in specific locations of the human genome. Scars can potentially happen anywhere at the whole genome level, and so genomic scarring interpretation requires NGS data that covers the entire genome.
The most promising genomic scarring approaches rely on high coverage whole genome sequencing (WGS) data and often require sequencing both the tumor sample and a normal tissue-matched control. While these approaches are effective, they are not suitable for routine testing in the clinical setting. The cost of high coverage WGS is still prohibitive and often sequencing paired tumor and control tissues is not feasible.
We hypothesized that WGS sequencing performed at low coverage (<1x) could contain sufficient information to detect HRD in samples. To test this hypothesis, we worked on publicly available datasets. These datasets contained high coverage matched tumor-normal WGS, for which the HRD status was established via genomic scars. We conceived a machine learning approach aimed at predicting HRD status by only looking at 1% of the original data in the tumor-only samples, still sequenced genome-wide but at a coverage of 1x or less. Surprisingly, we found that 1% of the data was sufficient to achieve excellent performance in predicting the HRD status that was previously established using 100% of the data.
In collaboration with the SOPHiA GENETICS genomic research laboratory, we explored the possibility of extending our library preparation and sequencing pipeline to generate a single workflow that includes: 1) the generation of low coverage WGS data for the detection of genomic scars, and 2) high coverage data for the targeted detection of mutations in HRR genes.
SOPHiA GENETICS’™ standard solutions rely on a targeted sequencing approach whereby an NGS-based WGS library is prepared, enriched, and sequenced to obtain high coverage NGS data in specific genomic regions of interest where variant calling will be performed. One might say that the technology already existed. The only modification was to load the sequencer with both the enriched library and a small amount of the WGS library (before enrichment). Thus, the SOPHiA DDM™ HRD Solution can simultaneously, and economically, generate WGS and targeted sequencing data to detect HRD via a genomic scarring approach (machine learning on WGS data), as well as via a traditional approach (variant calling on targeted data).
The SOPHiA DDM™ HRD Solution can simultaneously, and economically, generate WGS and targeted sequencing data to detect HRD.
Variant calling in HRR genes cannot entirely solve the HRD challenge. The Genomic Integrity Index is computed using our deep learning (a type of machine learning) approach and is a measure of a tumor that while being BRCA wild-type, is HRD positive. Our solution thus allows for the identification of additional HRD positive cancers that can benefit from PARPi.
Deep learning is particularly efficient in solving image classification tasks. A classical textbook example is, given an image, tell me if the image is of a cat or a dog. Convolutional neural networks solve this problem by learning from data which, in this example, is a dataset of images showing cats or dogs. The features present in the images that are best suited to distinguish dogs from cats are selected to train the data. After optimizing the parameters of the neural network, the convolutional neural network can take an image, that was not seen before, and determine whether it is of a cat or a dog.
Our technology works in the same way. The only difference is that the image used as input shows the coverage profile obtained from low coverage WGS data. The task of the convolutional neural network is to establish if the image comes from an HRD-positive or HRD-negative tumor sample.
After training and testing our deep learning algorithm on public data, we started working on clinical FFPE samples from ovarian cancer patients. This work was done in collaboration with some prestigious labs, including Diagnosticos da America SA (DASA) in Brazil. These samples were simultaneously tested with our HRD solution and gold standard methods used for HRD testing. The excellent concordance observed in these studies confirmed the validity of our solution.
Using a single workflow, our solution allows for the simultaneous generation of targeted sequencing and WGS data. Our deep learning algorithm allows us to make accurate predictions on the HRD status of a tumor sample, without requiring a matched normal sample, making the approach more feasible. Given the power of our deep learning algorithm, only a limited amount of WGS data is required. This allows for multiplexing up to 24 samples in a single Illumina NextSeq® 550, making the solution economically viable for routine testing.
Our deep learning algorithm allows us to make accurate predictions on the HRD status of a tumor sample.
The ESMO guidelines recommend the incorporation of HRD testing in addition to BRCA testing for women with newly diagnosed advanced ovarian cancer.2 Approved solutions for HRD tests fail to reliably identify patients who will not respond to PARPi. This is compounded by the fact that current solutions for HRD detection are expensive and sometimes infeasible (e.g. require paired tumor-normal samples and use of send-out services). Clinical researchers are thus looking for a cost-effective and reliable test to determine HRD status in ovarian cancers and identify those that could benefit from PARPi in the first-line or maintenance therapies. That is exactly what the SOPHiA DDM™ HRD Solution provides – it allows clinical researchers to independently conduct an accurate and comprehensive 2-in-1 HRD test in-house, in a cost-effective and time-saving manner.
HRD is a common feature of high-grade serous ovarian cancer (HGSOC) as well as breast, prostate, and pancreatic cancers. Homologous recombination repair (HRR) is a DNA damage repair system responsible for the repair of DNA double-strand breaks. HRD may be used as a potential predictive biomarker for poly (ADP-ribose) polymerase 1 (PARP1) inhibition therapy response, based on its predicated synthetic lethality in the context of HRD-positive cells. HRD testing provides an opportunity to identify larger populations of cancer patients who may benefit from treatment with PARP inhibitors (PARPi).
Cells respond to DNA damage by initiating multiple interconnected DNA repair pathways. HRR represents a central high-fidelity DNA damage repair system of DNA double-strand breaks. Functional defects in HRR are called HRD.1 The most known causes of HRD are loss of function mutations in genes involved in HRR including BRCA1, BRCA2, RAD51C, RAD51D, PALB2, as well as promoter hypermethylation of BRCA1.2 Over time, unrepaired or inaccurately repaired double-strand breaks lead to the accumulation of genomic aberrations, such as insertions and deletions, copy number alterations, or structural chromosomal rearrangements. These manifest as genomic instability, which drives carcinogenesis and disease progression.1
HRD is identifiable in many cancers. Approximately 15% of HGSOC harbor germline BRCA1/2 (gBRCA1/2) mutations. A total of 50% have HRD, conferred by either germline or somatic mutations in BRCA1/2 or mutations in other genes known to be involved in HRR.3 Similarly, gBRCA1/2 mutations are harbored by approximately 15% of triple negative breast cancers (TNBC) and a further 40% have HRD in the absence of gBRCA1/2 mutations. In advanced prostate cancers, 10%–12% of patients have germline or somatic BRCA2 inactivation and up to 25% contain a DNA damage repair defect.4 It remains to be determined whether HRD in different cancer types has the same biological and clinical implications.2
HRD testing provides an opportunity to identify larger populations of cancer patients who may benefit from treatment with PARP inhibitors (PARPi)
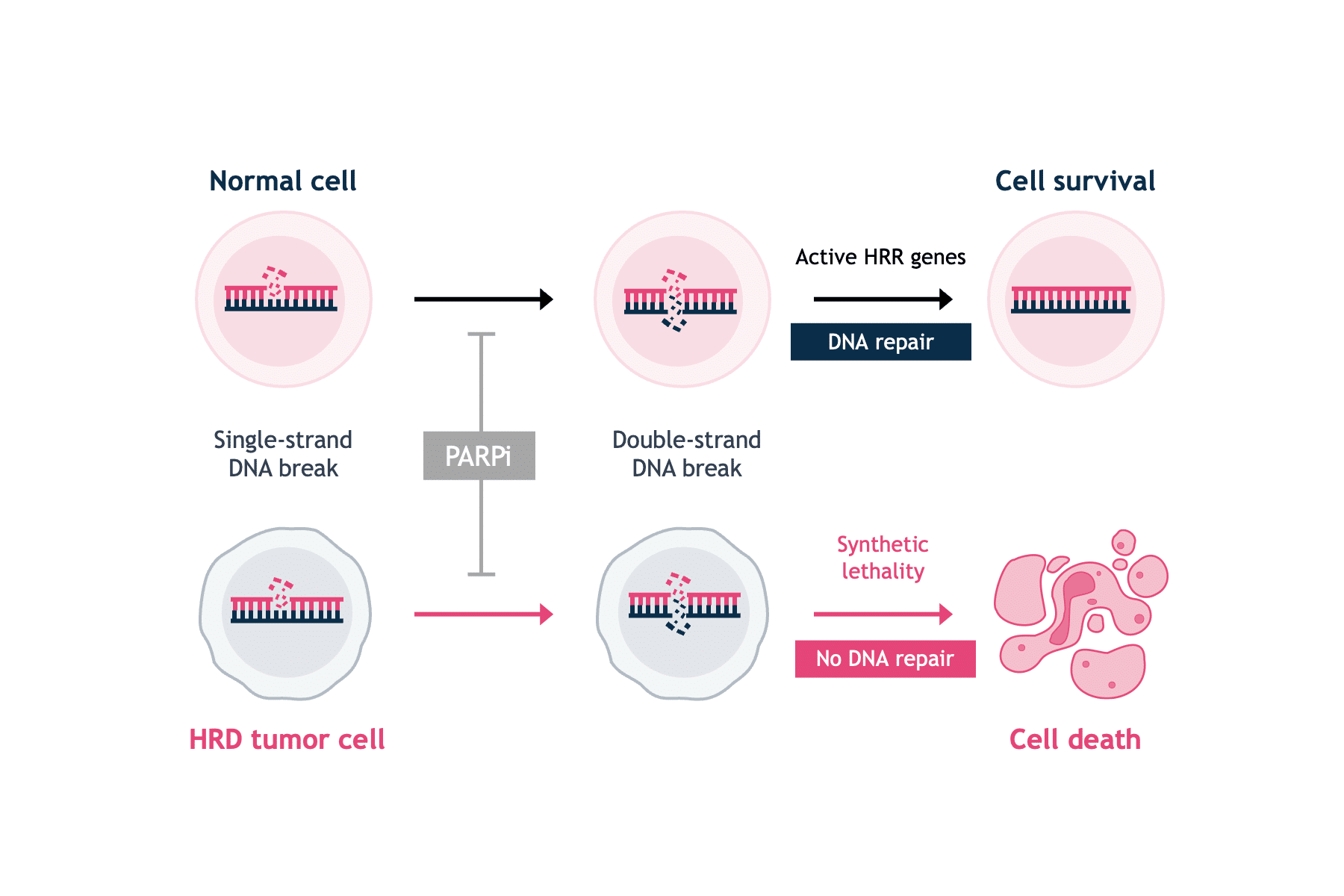
PARPi are small molecule inhibitors of the PARP family of proteins.5 PARP1 is best known for its role in the base excision repair of single-strand DNA breaks.1 The discovery that single-agent PARP inhibition selectively killed BRCA-deficient cells was a breakthrough in exploiting synthetic lethality approaches in oncology (Figure 1).4,6
Pre-clinical evidence has shown that PARP can modulate the repair of double-strand DNA breaks, if cancer cells are HRD without the capacity to repair double-strand DNA breaks.7 PARPi trap the PARP1 protein on DNA at sites of single-strand DNA breaks. If trapped PARP1 is encountered by the DNA replication machinery, it leads to stalling of the replication fork, the collapse and the generation of a double-strand break, which cannot be repaired in HRD cells. Thus, PARPi prevent the DNA repair process, ultimately leading to the disruption of cellular homeostasis and cell death.8 HRD has been identified as predictive biomarker for PARPi therapy response in ovarian, and potentially in breast, and prostate cancers.9–11
The assessment of HRD is an important prognostic and predictive biomarker.1 Currently available testing methods for HRD fall into three categories: HRR gene-based tests, genomic scars and signatures, and functional assays.4
HRR gene-based tests
The most reliable clinical predictor of HRD and response to PARPi remains gBRCA mutations and FDA approval has been granted for gBRCA detection tests in ovarian cancer.12 However, the negative predictive value of BRCA mutation status is poor in the setting of platinum-sensitive relapsed HGSOC, with BRCA wild-type (BRCAwt) subgroups also deriving a significant, although numerically smaller benefit from PARPi. There are currently no approved diagnostic tests for HRD based on germline mutations of other HRR genes.4 Using germline mutations in HRR genes to predict HRD status has several drawbacks. Genetic testing often results in the detection of variants of uncertain significance where a particular rare genotype may not confer an HRD phenotype. Additionally, somatic reversion mutations in the BRCA genes can restore homologous recombination proficiency (HRP). HRDstatus is therefore a complex and dynamic phenotype.4
Genomic scars and signatures
Since a considerable number of genes with potentially variable biological impact are involved in HRR, elucidating the clinical implications of individual germline or somatic mutations in HRR coding genes is challenging. An alternative strategy is to identify the effects of deleterious mutations in HRR genes. The inability to repair DNA in tumors that have HRD results in genomic scars or mutational signatures that measure the patterns of somatic mutations that accumulate as a result of HRD, irrespective of the exact nature of the underlying etiology.1,4 Studies of genomic scars and signatures generally measure somatic copy number variation (CNV) and predict BRCA status through the quantification of large-scale transitions (LST), loss of heterozygosity (LOH) or the number of sub-chromosomal regions with allelic imbalance extending to the telomere (NtAI), combined in most tests to generate a genomic instability score.4 Current commercially available tests combine tumor BRCA mutation testing with a genomic instability score.
Functional assays
Conversely, functional HRD assays have the potential to provide a dynamic readout of the current HRR status. The most commonly used experimental system to estimate HRR has been to assess the amount of nuclear RAD51.4 RAD51 is a downstream protein in the HRRpathway that is loaded onto sites of DNA double-strand breaks to facilitate DNA strand invasion into the sister chromatid in cooperation with mediator protein complexes, especially BRCA1 and BRCA2 (Figure 2).1,4

HRD cells are characterized by a reduction or inability to form RAD51 foci. This phenotype occurs irrespective of the underlying cause of HRD. In the presence of acquired reversion mutations that restore HRR, RAD51 foci are present. However, this technique also has its caveats. The RAD51 assays will not identify HRR defects downstream of RAD51 loading on to DNA and the RAD51 signal may be elicited by exogenous DNA damage. Another functional measure of HRD is the sensitivity to platinum salts, which in itself is a predictive marker for PARPi.5 Platinum sensitivity is a feature of HRD cells, and BRCA-mutant ovarian and breast tumors demonstrate increased platinum sensitivity.12 Since platinum-based therapies are a key component of first-line chemotherapy in ovarian cancer, prior platinum sensitivity has been evaluated as a surrogate clinical index for prediction of efficacy to PARP inhibition.12 However, there is still insufficient evidence to ascertain the clinical validity of functional assays in predicting PARP inhibition response.4
At SOPHiA GENETICS, we offer a innovative approach to HRD detection with the SOPHiA DDMTM HRD Solution. In a single application, our solution allows clinician researchers to:
The comprehensive application provides researchers with timely, in-house results on HRD status.
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™ Dx Hereditary Cancer Solution, SOPHiA DDM™ Dx RNAtarget Oncology Solution and SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution are available as CE-IVD products for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA DDM™ Dx Myeloid Solution and SOPHiA DDM™ Dx Solid Tumor Solution are available as CE-IVD products for In Vitro Diagnostic Use in the EEA, the United Kingdom, Switzerland, and Israel. Information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us to obtain the appropriate product information for your country of residence.
All third-party trademarks listed by SOPHiA GENETICS remain the property of their respective owners. Unless specifically identified as such, SOPHiA GENETICS’ use of third-party trademarks does not indicate any relationship, sponsorship, or endorsement between SOPHiA GENETICS and the owners of these trademarks. Any references by SOPHiA GENETICS to third-party trademarks is to identify the corresponding third-party goods and/or services and shall be considered nominative fair use under the trademark law.