Menu
Menu
Assessment of Homologous Recombination Deficiency (HRD) and BRCA mutational status in high-grade serous ovarian carcinoma (HGSOC) samples has become crucial for personalized medicine, guiding treatment decisions, and predicting response to specific therapies, such as PARP inhibitors.
The aim of this multicentric study, published at this year’s European Society of Gynaecological Oncology (ESGO) conference was to assess the reliability and reproducibility of SOPHIA DDM™ Dx HRD Solution across three different hospitals (Hospital del Mar, Hospital Vall d’Hebrón and Hospital Clinic de Barcelona) to consider its implementation as a decentralized solution in routine molecular diagnostics.
Our Spanish team led by Giovanni Velotta, CS Manager at SOPHiA GENETICS, was happy to meet Dr. Gardenia Vargas, Molecular Geneticist and Responsible for Hereditary Cancer and Rare Disease Molecular Diagnosis at Hospital del Mar in Barcelona, Spain, who joined us for an insightful interview, sharing her experience in implementing SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution.
Before I answer I would like to thank you and thank AstraZeneca for their support in this study. I must mention that all my answers represent my own perspectives and not necessarily the official stance of the hospitals involved in this project.
The study aimed to investigate the feasibility and effectiveness of implementing the SOPHiA DDM™ Dx HRD (Homologous Recombination Deficiency) Solution within the clinical setting of three hospitals. This includes evaluating various aspects such as the practicality of integrating the test into existing diagnostic workflows, the accuracy and reliability of test results, the impact on patient care and outcomes, and the potential benefits and challenges associated with in-house testing. The study seeks to provide valuable insights into the utility and suitability of adopting the SOPHiA DDM™ Dx HRD (Homologous Recombination Deficiency) Solution as a routine diagnostic tool for identifying HRD in ovarian cancer patients.
First of all, I must say that all three hospitals worked equally on this project, and the idea of doing it was born before I was incorporated to it. Beatriz Bellosillo from Hospital del Mar was the principal investigator, and we had different responsibilities such as protocol writing, ethics approval coordination, securing funding to support the study's activities, and management and allocation of some resources like sequencing supplies.
And of course, it was truly a rewarding experience collaborating with renowned hospitals and esteemed colleagues.
Currently, while we have conducted a multicenter study to evaluate the feasibility of implementing the SOPHiA DDM™ Dx HRD Solution, we have not yet implemented it into routine diagnostic practice at my hospital.
The implementation process requires significant financial support, particularly for acquiring the necessary equipment, reagents, and personnel training. Additionally, to ensure cost-efficiency and timely responses to patients, we need to conduct sequencing runs with a minimum number of samples per run.
Unfortunately, our hospital alone does not have sufficient sample volume to meet this requirement. Therefore, we are actively seeking financial support from government agencies to fund the implementation of the study. Additionally, we are exploring collaborative efforts with other hospitals or healthcare institutions to pool together an adequate number of samples for sequencing runs, thus reducing costs per sample and ensuring a rapid response to patients.
By securing both financial support and an adequate sample volume, we aim to overcome these logistical challenges and proceed with the implementation of the SOPHiA DDM™ Dx HRD Solution into our diagnostic routine, ultimately enhancing our ability to provide efficient and effective patient care.
It highlights the importance of collaboration among hospitals, government agencies, and other stakeholders to support the implementation of advanced diagnostic technologies into routine clinical practice.
Overall, the study provides valuable insights into the challenges and opportunities associated with integrating advanced genomic testing into routine diagnostics, with the aim of improving patient care and outcomes.
This collaborative study concludes that SOPHIA DDM™ Dx HRD Solution provides reliable and consistent results across different hospitals and sequencing runs.
These findings contribute to the growing body of evidence supporting the use of the decentralized SOPHiA DDM™ Dx HRD Solution in clinical settings for genomic analysis.
DISCLAIMER: SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise. SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution is available as CE-IVD product for In Vitro Diagnostic Use in Europe and Turkey.
Explore this infographic summary to learn more about the machine learning model developed by Margue et al. for the prediction of disease-free survival in patients undergoing surgery for renal cell carcinoma.

Margue G, et al. UroPredict: Machine learning model on real-world data for prediction of kidney cancer recurrence (UroCCR-120). NPJ Precis Oncol. 2024 Feb 23;8(1):45.
SOPHiA GENETICS multimodal analysis technology and concepts in development. May not be available for sale.
Explore this infographic summary to gain insights into the key findings from Pozzorini et al.’s publication on the GIIngerTM deep learning algorithm for prediction of HRD status and patient response to PARPi treatment in ovarian cancer.
Click here to read the full publication.
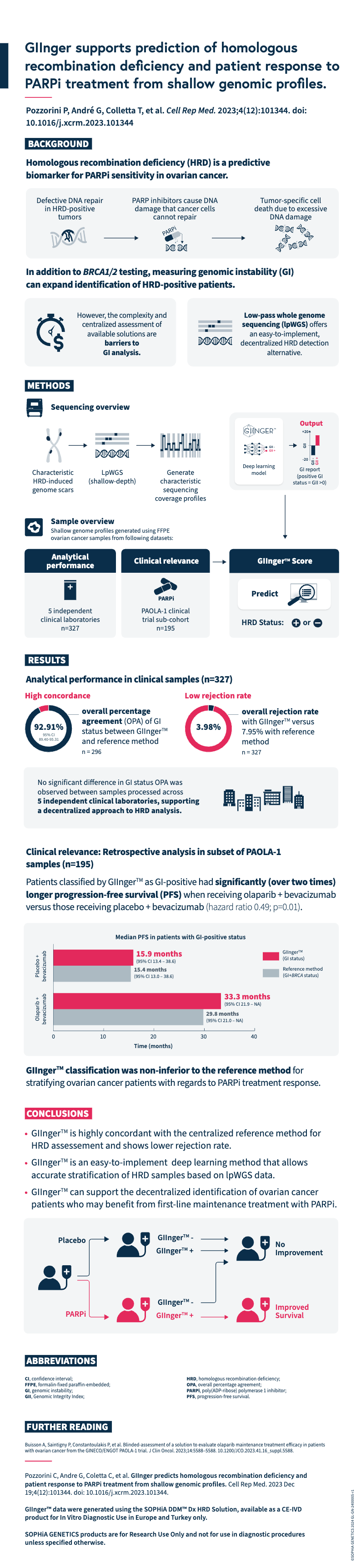
Pozzorini C, Andre G, Coletta C, et al. GIInger predicts homologous recombination deficiency and patient response to PARPi treatment from shallow genomic profiles. Cell Rep Med. 2023 Dec 19;4(12):101344. doi: 10.1016/j.xcrm.2023.101344.
GIInger™ data were generated using the SOPHiA DDM™ Dx HRD Solution, available as a CE-IVD product for In Vitro Diagnostic Use in European Economic Area (EEA), the United Kingdom and Switzerland . SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
Do you want to further your understanding of overfitting, or are you interested to learn how we avoid overfitting when developing predictive machine learning models for clinical research? Browse our tech note, which includes a step-by-step, published example of how we developed a model that was able to successfully support the evaluation of kidney cancer tumor upstaging in individual patients.
Genomic instability is a hallmark of cancer and targeting its mechanisms has helped inform effective therapeutic strategies1,2. However, there are limitations with current methods of genomic instability assessment. Here, we explore genomic instability in the context of homologous recombination deficiency and the value of deep learning-based methods of detection.
The DNA in our cells endure up to one million damaging events each day, caused by both exogenous (i.e. environmental) and endogenous (i.e. internal metabolic) factors3. These events activate a complex network of DNA damage response (DDR) pathways, which facilitate DNA repair and maintain the stability of the genome4. Mutations and/or dysfunction in DDR pathways can lead to unrepaired DNA damage, resulting in genomic instability4. Genomic instability, in turn, increases the cell’s propensity for genetic alterations that cause cancer initiation and progression4,5.
One of the major DDR pathways is the homologous recombination repair (HRR) pathway, responsible for repairing double-strand breaks (DSBs) in DNA5. Loss of function in HRR, known as homologous recombination deficiency (HRD), causes cells to rely on error-prone DNA repair pathways, resulting in the accumulation of genetic aberrations that lead to genomic instability5 (Fig 1). HRD is a well-established prognostic and predictive biomarker in different cancer types (e.g. ovarian, breast, prostate, and pancreatic)5–7.
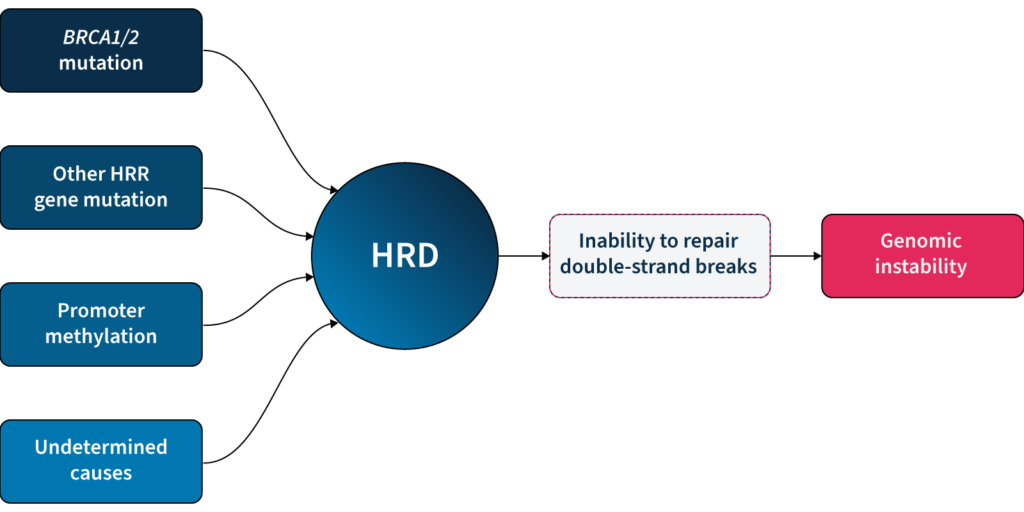
HRD-positive tumors are sensitive to targeted inhibition of poly-ADP ribose polymerase (PARP), key proteins involved in DSB repair7. By blocking PARP, the HRD-positive cell can no longer rely on error-prone pathways for DSB repair and the cell dies, a process known as ‘synthetic lethality’5,6. PARPi therapy has revolutionized the management of HRD-positive patients with advanced ovarian cancer, significantly improving progression-free survival when used as a first-line maintenance therapy8. PARPi therapies also have approved indications in breast, pancreatic and prostate cancer9, with trials underway in other cancer types, such as colorectal10.
Based on the predictive value of HRD status for PARPi benefit, clinical guidelines recommend HRD testing in patients with advanced ovarian cancer7,11,15. HRD status can be determined by examining 1) the underlying causes of HRD, and 2) the effect of HRD, i.e. genomic instability5,7. The most well-known causes of HRD are loss-of-function mutations in HRR genes, including BRCA1 and BRCA25,7. However, loss-of-function in HRR genes is diverse amongst patients12, making patient stratification solely based on genotyping challenging. Also, approximately 30–40% of HRD cases are due to unknown causes13,14. Measuring genomic instability allows the assessment of HRD, regardless of its underlying etiology5,7.
Genomic instability status can help identify a sub-group of women who are BRCA wild-type but may still derive benefit from PARPi therapy15. By measuring genomic instability, clinicians and researchers can therefore go beyond HRR mutation detection and expand the potential benefit of PARPi in patients.
Many methods for measuring genomic instability rely on the identification of specific mutational signatures or genomic ‘scars’ associated with large-scale structural rearrangements in chromosomes. In HRD-positive cancers, the characteristic genomic scars are loss of heterozygosity (LOH), large-scale state transitions (LST), and telomeric-allelic imbalance (TAI)16–18.
Click the boxes below to learn more:

A cross-chromosomal event that results in loss of part of a gene or entire gene(s) and the surrounding chromosomal region.

Chromosomal breaks between adjacent regions of at least 10 Mb.
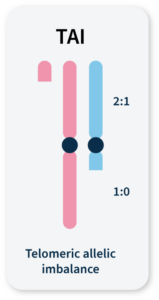
Accumulation of a discrepancy in the 1:1 allele ratio at the end of the chromosome (telomere).
The combined number of LOH, LST, and TAI events generate a genomic instability score (GIS) that reflects the level of genomic instability. Some commercially available HRD tests combine tumor BRCA mutation testing with a GIS5,19. Methods that integrate multiple genome-wide signatures (e.g. HRDetect) are among the most promising for detecting HRD status7,20. However, both GIS and HRDetect methods require deep genomic profiling data (>30x coverage), which can be costly and difficult to implement in routine analysis.
Alternative approaches that rely on the detection of copy number changes from WGS at low (~1x) sequencing depth (low-pass WGS) can predict tumor HRD status21,22 and provide an affordable and easy-to-implement HRD detection method. However, the sensitivity of existing methods that solely rely on this type of genomic scar to identify HRD samples is limited, and their utility in a clinical context remains untested22.
Unlocking the full potential of low-pass WGS in HRD detection requires going beyond the enumeration of biomarker events and examining alternative features of the cell that can result from genomic instability.
To overcome the limitations of current genomic instability measures, our expert team at SOPHiA GENETICS developed the GIInger™ algorithm exclusively available on the SOPHiA DDM™ Platform. GIInger™ is a deep learning-based approach to measuring genomic instability in ovarian cancer samples. Rather than relying on the enumeration of biomarkers events, GIInger™ leverages differences in the spatial distribution of genomic scars in low-pass WGS coverage profiles23.
Let’s take a closer look at how the algorithm predicts genomic instability status (Fig 2):
Unfamiliar with deep learning terminology? Read our guide on machine learning jargon.
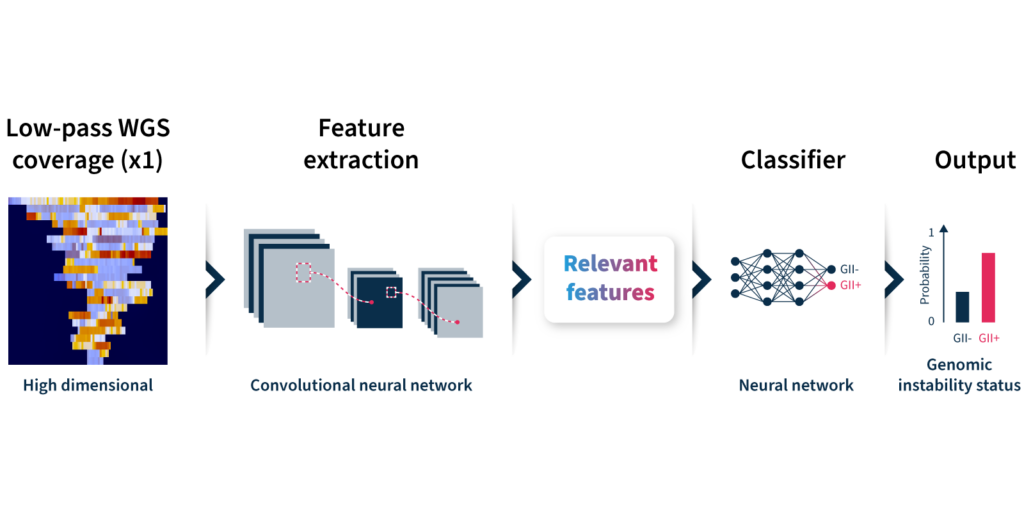
By adopting GIInger™ into next generation sequencing (NGS) workflows, clinical researchers can benefit from an in-house, affordable approach to genomic instability measurement. The SOPHiA DDM™ Platform offers applications that enable laboratories to easily implement GIInger™ into their routine NGS analysis:
Want to see how GIInger™ can help maximize insights from your data? Get in touch with our team and request a demo.
The term « SOPHIA » used by the speaker refers to SOPHiA GENETICS and its products. The opinions expressed during this presentation are these of the speaker and may not represent the opinions of SOPHiA GENETICS. SOPHiA GENETICS does not provide support in the validation of custom products for clinical use. SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution is available as a CE-IVD product for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™️ Dx ROS is a novel CE-IVD marked NGS assay for detecting gene fusion and exon skipping events from small lung cancer biopsy samples. A performance evaluation study demonstrates that, compared with existing RNA-based assays, SOPHiA DDM™️ Dx ROS provides accurate actionable clinical insights for healthcare professionals to make disease management decisions for patients with advanced lung cancer
At ESMO 2022, oncology experts gathered in Paris and online to share and debate the new developments in the field of medical oncology. This year's program featured more than 20 tracks covering all tumor types, therapeutic innovations, translational research, patient advocacy, public policy, and more...
Discover our summary of three compelling talks showcasing the journey towards precision therapy in various tumor types.
Genomic profiling and molecular targeting of lung cancer brain metastases1
Haiying Cheng, Dept. Medical Oncology, Montefiore Medical Center/Albert Einstein College of Medicine, Bronx, NY, USA
Approximately 57% of patients with non–small-cell lung cancer (NSCLC) present with metastatic disease2. Among them, brain metastases (BM) affect up to 45% of all cancer patients and arise from lung cancers in 40-50% of the cases3. There have been limited studies investigating the genetic signatures of LC BM, and with small cohorts so far.
Assembling a large number of lung cancer cases (47215 NSCLC; 29438 lung adenocarcinoma), Dr Cheng and colleagues looked for key genetic alterations in loco-regional lesions (Loco), extracranial metastases (EM), and BM with comprehensive genomic profiling (CGP). They found significantly more genetic alterations in the PI3K/AKT/mTOR pathway in BM (Loco 13.0% vs EM 14.5% vs BM 18.1%), primarily driven by RICTOR amplification (Loco 3.6% vs EM 6.2% vs BM 8.6%).
RICTOR amplification is the most enriched actionable genomic target in NSCLC brain metastases.
Furthermore, in vitro genetic knockdown and pharmacological inhibition of RICTOR significantly reduced migration and invasion in RICTOR-amplified NSCLC cells, whereas RICTOR upregulation promoted these processes, modulating the AKT, MET, EMT, and CXCL12 chemokine-CXCR4 pathways. Finally, in vivo studies in orthotopic mouse models revealed that both RICTOR and mTOR1/2 inhibition significantly reduced lung cancer tumor growth and spread in the brain.
Dr Cheng provided evidence for the benefit of further investigation on the development of RICTOR-targeted therapeutic strategies for the treatment and/or prevention of lung cancer BM. This study is a good example of how genomic profiling, combined with functional analyses, can identify new potential therapeutic targets.
Neoadjuvant immune checkpoint inhibition in locally advanced MMR-deficient colon cancer: The NICHE-2 study4.
Myriam Chalabi, Gastrointestinal Oncology, Netherlands Cancer Institute, Amsterdam, Netherlands
Mismatch repair deficiency (dMMR) is observed in ~15% of colorectal cancers (CC)5 and 1/3 is associated with Lynch Syndrome. This characteristic genetic signature is marked by high levels of microsatellite instability (MSI) and resistance to standard-of-care neoadjuvant chemotherapy (5-7% pathological response (≤50% residual viable tumor; PR)). NICHE-1 exploratory study showed the potential of neoadjuvant immunotherapy in patients with dMMR CC with extraordinary PR in 100% of the patients6.
Dr Chalabi presented the NICHE-2 investigator-initiated study, conducted in 6 hospitals in the Netherlands. 107 patients with non-metastatic untreated dMMR CC and mainly high-risk tumors received injections of nivolumab and ipilimumab within 6 weeks prior surgery. The impressive pathological tumor regression was shown in a waterfall plot that led to a standing ovation! 95% of the treated patients showed PR, and 67% had no residual viable tumor (complete PR; cPR), contrasting with the neoadjuvant chemotherapies in the same patient population. Only 4% experienced grade 3-4 immune-related adverse events and 98% of patients underwent timely surgery, meeting the safety primary endpoint. To date, no disease recurrence has been observed and the 3 years disease-free survival data are expected next year.
Neoadjuvant immunotherapy has the potential to become standard of care for patients with dMMR colon cancer.
NICHE-2 trial opens the possibility that a surveillance approach may be possible for some patients with early dMMR CC and gives a glimpse at the potential of translational research to identify predictive biomarkers in pre- and post-treatment samples. While those preliminary results are extremely promising and we surely wait for the longer-term follow-up data to confirm them, patient selection remains crucial. Indeed, neoadjuvant decisions are based on radiological assessment of the tumor, particularly difficult in dMMR cancers, as well as the biopsy, and Dr Chalabi highlighted the need for improvement in the imaging techniques and circulating DNA analyses.
Final overall survival results from the phase III PAOLA-1/ENGOT-ov25 trial evaluating maintenance olaparib plus bevacizumab in patients with newly diagnosed advanced ovarian cancer7.
Isabelle Ray-Coquard, Department Of Medical Oncology, Centre Léon Bérard, and GINECO, Lyon, France
The late diagnosis of advanced ovarian cancer (AOC) is often accompanied by relapse, despite surgery and platinum-based chemotherapy. Treatment with olaparib (ola), a poly(adenosine diphosphate–ribose) polymerase inhibitor (PARPi), provided progression-free survival (PFS) benefit as maintenance therapy in patients with AOC carrying mutations in BRCA1 or BRCA2 (BRCAm)8. Besides, the incorporation of the antiangiogenic agent bevacizumab (bev) is a recognized option in addition to chemotherapies9.
PAOLA-1 investigators conducted a phase III trial where 806 patients with AOC and after first-line platinum-based chemotherapy plus bev were randomly assigned in a 2:1 ratio to ola + bev or placebo (pbo) + bev treatment. The primary endpoint was the PFS. They showed that combined treatment with ola + bev reduced the risk of relapse by 41% compared to bev alone, reducing by 67% in HRD+ patients (exhibiting BRCAm and/or genomic instability score ≥42)10.
Here, Dr Ray-Coquard presented the final overall survival (OS) results, a key secondary endpoint. She showed that the OS rate after 5 years was not different between the two arms (47.3% vs 41.5%) but significantly increased for HRD+ patients treated with ola + bev (65.5% vs 48.4%), regardless of BRCAm status. Also, PFS was significantly increased in the same population (46.1% vs 19.2%).
Maintenance therapy with olaparib plus bevacizumab improved survival in HRD+ patients with newly diagnosed advanced ovarian cancer.
With the absence of new safety signals and major adverse effects, these data confirmed the benefit of olaparib and bevacizumab combination as a standard of care for HRD+ patients and reinforced the importance of precision medicine and biomarker testing to guide treatment decisions.
Meet the new SOPHiA DDM™ Dx RNAtarget Oncology Solution (ROS).
Cancer management outcomes can strongly benefit from robust and accurate detection of gene fusions. SOPHiA GENETICS developed the SOPHiA DDM™ Dx RNAtarget Oncology Solution (ROS) to make this possible.
Gene fusions are “hybrid genes” formed by the fusion of two genes into one and producing a novel protein. These are mostly due to DNA translocations, but also sometimes resulting from DNA insertions, inversions, duplications, or deletions1. Exon-skipping events, the erroneous excision of one or more exons during RNA splicing, also lead to pathologically structured genes. Both play an important role in carcinogenesis1 and, because of this, have been preferential targets of precision medicine as well as of some of the most effective treatments in fusion-positive cancers2,3.
Whether specific rearrangements occur at DNA or RNA level, they can both lead to new proteins with altered functions: in human cancers, more than 10,000 gene fusions creating oncogenic drivers of specific tumors or activators of oncogenic signalling have been identified4. Given the large number of gene fusions to identify, researchers worldwide increasingly need intelligent bioinformatics tools to automate the process - and even more: detect them with suitable technical benefits, like the possibility of customizing gene panels, the use of a minimum input sample, and the chance to optimize the lab workflow using high sensitivity technology. Considering the importance of gene fusions as a diagnostic and prognostic marker for cancer management in the clinical setting5-7, these should be transitioned from the research to patient's bedside. In this particular context, the precious RNA sample for gene fusion testing is not only a “simple sample” - it represents a unique and individual opportunity for better treatment outcomes8. The right technology choice sets the tone for high-accurate detection of gene fusions and, consequently, for high-precision medicine.
Moreover, the possible association between gene fusion type and tumor phenotype can flag the clinical value of accurate gene fusion detection for:
The right technology choice sets the tone for high-accurate detection of gene fusions and, consequently, for high-precision medicine.
Multiple approaches have been developed to effectively detect gene fusions at DNA, RNA, and protein level based on their significant clinical relevance. The investigation of gene fusions at the DNA level allows to identify structural chromosomal rearrangements, while looking at RNA uncovers fusions generated during RNA splicing without any chromosomal rearrangement and potentially translated into defective fusion proteins. Gene fusion identification has been traditionally performed by immunohistochemistry, fluorescence in situ hybridization (FISH), or polymerase chain reaction (PCR), but next-generation sequencing (NGS) methods represent the current gold standard6,7. These high-throughput sequencing technologies can identify gene fusions either at DNA or RNA level. NGS has become the technology of choice for fusion detection in solid tumors in agreement with international guidelines7. The simultaneous analysis of detailed (nucleotide-level) and comprehensive (genome-wide) information on the genome or transcriptome (all RNA transcripts) allow searching for the constantly increasing number of fusion genes, which would be otherwise impossible to achieve without high-throughput techniques8,9.
Although DNA-based sequencing technologies perform a comprehensive investigation, their short-read NGS approach currently hinders the proper mapping of several gene fusions. Furthermore, they require a considerable amount of sequencing, storage space, and long computational analysis, which makes them expensive and time-consuming. On the other hand, RNA-based approaches are more cost-effective: less storage space and analysis time are needed since only genetic regions which are transcribed and spliced into mature mRNA are explored; therefore, expressed gene fusions are exclusively detected. Also, RNA-based approaches overcome possible technical limitations of DNA-based methods derived from the localization of fusion breakpoints either within long DNA intronic regions (difficult to capture) or on different sites on DNA and RNA encoding the same fusion because of post-transcriptional and splicing rearrangements.
The success of RNA-based gene fusion detection approaches relies on sample quality and quantity: proper RNA extraction, stability, abundance of mRNA transcribed from the gene of interest, and a sufficient amount of sample can often be challenging9. Technics using small inputs coming from precious biopsy samples are required. Furthermore, algorithms with high gene fusion prediction accuracy are crucial. Finally, the complexity of the analysis, which is computationally demanding, remains a significant challenge10.
SOPHiA GENETICS aimed to solve the current challenges of RNA-based approaches for gene fusion detection by developing the SOPHiA DDM™ Dx RNAtarget Oncology Solution (ROS).
Here we list seven reasons11 why you should consider SOPHiA DDM™ Dx ROS for your assays:
Compared to a fully-guided amplicon-based approaches, the capture-based design of SOPHiA DDM™ Dx ROS detects fusion without prior knowledge of the partners, increasing considerably the power of fusion detection and exon-skipping events.
SOPHiA DDM™ Dx ROS requires only 10 ng of RNA from formalin-fixed paraffin-embedded (FFPE) or fresh frozen tissues. No fragmentation is required before FFPE samples processing, while a short simple fragmentation is performed on fresh frozen tissues. The solution is also compatible with total nucleic acid (TNA).
Together with gene fusions, SOPHiA DDM™ Dx ROS can detect single nucleotide variants (SNVs) and insertion/deletion mutations (Indels). The SNV/Indels calling performed on RNA indicates expression and, therefore, the related biological relevance of the detected mutations is also provided.
The analytical requirements of SOPHiA DDM™ Dx ROS have been adapted to balance the solution's sensitivity and specificity. SOPHiA DDM™ Dx ROS variant caller identifies PCR duplicates to enhance results accuracy. The analytical requirements were optimized to balance the sensitivity and specificity of our technology. The analytical parameters were carefully chosen to balance the need to remove deamination artifacts and background noise. Furthermore, SNV calling algorithms account for biases due to the low biological expression level of the gene of interest. To achieve global standard results with high sensitivity detection of gene fusions, SOPHiA DDM™ Dx ROS was designed based on the recent ESMO (Tier I and II) and NCCN international guidelines for lung cancer6. The probabilistic model applied to the data leverages multiple features to deliver results with reduced false positives and high accuracy of fusion calling.
SOPHiA DDM™ Dx ROS is easily adaptable to the unique needs of your laboratory. An optimized workflow with a dedicated probe design process and extensive wet lab QC experiments provides you with a ready-to-use panel, reducing the need for additional optimization in the lab.
SOPHiA DDM™ Dx ROS comes with a streamlined 1.5-day workflow that supports the whole analysis and ensures access to record-time results.
The SOPHiA DDM™ Platform fully supports you in the data analysis and interpretation process: pipelines are adapted to sample type, sequencer, enrichment kit, and chemistry used in your laboratory. Moreover, s streamlined workflow helps you to annotate, interpret, and report relevant NGS variants. The complexity of the analysis is minimized. The intuitive and user-friendly interface as well as its accelerated data visualization and interpretation and its customizable reporting and comprehensive QA report increase the efficiency of the decision-making process. The OncoPortal™ Plus module, gathering the latest scientific evidence and guidelines on the actionability and significance of each genomic alteration, completes the all-around support of the SOPHiA DDM™ Platform to facilitate your interpretation process.
SOPHiA DDM™ Dx ROS supports your laboratory in gene fusion detection. With its high sensitivity and accuracy, it allows the identification and the expression assessment of novel gene fusions, SNVs and Indels. It can also adapt to your research needs thanks to a fully customizable gene panel. Moreover, a streamlined workflow and the SOPHiA DDM™ Platform support you throughout the whole assay, from the RNA sequencing to the data analysis and interpretation. With SOPHiA DDM™ Dx ROS, you will maximize the insights from precious biopsy specimens getting highly reliable results even from samples of limited quantity.
1. Mitelman F, Johansson B, Mertens F (eds.) (2016) Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer,.http://cgap.nci.nih.gov/Chromosomes/Mitelman.
2. Taniue K, Akimitsu N. Fusion Genes and RNAs in Cancer Development. Noncoding RNA. 2021 Feb 4;7(1):10.
3. Brien GL, Stegmaier K, Armstrong SA. Targeting chromatin complexes in fusion protein-driven malignancies. Nat Rev Cancer. 2019 May;19(5):255-269.
4. Bruno R, Fontanini G. Next Generation Sequencing for Gene Fusion Analysis in Lung Cancer: A Literature Review. Diagnostics (Basel). 2020 Jul 27;10(8):521.
5. Mertens F, Johansson B, Fioretos T, Mitelman F. The emerging complexity of gene fusions in cancer. Nat Rev Cancer. 2015 Jun;15(6):371-81.
6. National Comprehensive Cancer Network®. NCCN Clinical Practice Guidelines in Oncology. Non-Small cell Lung Cancer. Version 3.2022
7. Mosele F, et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: a report from the ESMO Precision Medicine Working Group. Ann Oncol. 2020 Nov;31(11):1491-1505.
8. Ozsolak F, Milos PM. RNA sequencing: advances, challenges and opportunities. Nat Rev Genet. 2011 Feb;12(2):87-98.
9. Kaya C, et al. Limitations of Detecting Genetic Variants from the RNA Sequencing Data in Tissue and Fine-Needle Aspiration Samples. Thyroid. 2021 Apr;31(4):589-595.
10. Uhrig S, et al. Accurate and efficient detection of gene fusions from RNA sequencing data. Genome Res. 2021 Mar;31(3):448-460.
11. Data on file
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™ Dx Hereditary Cancer Solution, SOPHiA DDM™ Dx RNAtarget Oncology Solution and SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution are available as CE-IVD products for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA DDM™ Dx Myeloid Solution and SOPHiA DDM™ Dx Solid Tumor Solution are available as CE-IVD products for In Vitro Diagnostic Use in the EEA, the United Kingdom, Switzerland, and Israel. Information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us to obtain the appropriate product information for your country of residence.
All third-party trademarks listed by SOPHiA GENETICS remain the property of their respective owners. Unless specifically identified as such, SOPHiA GENETICS’ use of third-party trademarks does not indicate any relationship, sponsorship, or endorsement between SOPHiA GENETICS and the owners of these trademarks. Any references by SOPHiA GENETICS to third-party trademarks is to identify the corresponding third-party goods and/or services and shall be considered nominative fair use under the trademark law.