Menu
Menu
Precision oncology is transforming cancer care by tailoring treatment decisions to the unique molecular profile of each patient’s disease. Central to this approach is biomarker testing – however, no single test can capture the full biological complexity of cancer. Tissue biopsy and liquid biopsy can be complementary tools that answer different but equally important clinical questions, achieving a complete and dynamic molecular picture.
In this blog, we will explore insights from the ROME trial, which highlight the potential value of integrating both biopsy modalities for solid tumor analysis1.
Why two biopsies may be better than one
Tissue biopsy remains the cornerstone of cancer diagnostics, providing histology, tumor grading, and a deep view of local genomic alterations within a specific lesion. It is essential to confirm tumor type and assess immuno-oncology biomarkers such as microsatellite instability (MSI) and tumor mutational burden (TMB). However, tissue sampling is invasive, may not be repeatable, and captures only a snapshot from one tumor region at a single time point, potentially missing spatial and temporal heterogeneity2.
Liquid biopsy, by contrast, uses circulating tumor DNA to give a minimally invasive, real‑time snapshot of the genomic landscape across multiple tumor sites. It enables dynamic monitoring of treatment response and resistance mechanisms and can better reflect metastatic disease2, but may miss alterations in low-shedding tumors and does not provide histologic context3.
Together, the complementary strengths of both approaches may offer a combined strategy that provides a more complete molecular picture than either approach alone.
Inside the ROME trial design
The ROME trial was a phase II, multicenter, randomized study that enrolled 1,794 adults with advanced solid tumors in second or third line of therapy, regardless of histology. All patients underwent next‑generation sequencing on both tumor tissue and plasma with results reviewed by a centralized molecular tumor board (MTB). When actionable alterations were identified, 400 patients were randomized 1:1 to receive MTB‑guided tailored therapy (TT; targeted or immunotherapy) or standard-of-care (SoC) selected by the treating clinician, with crossover allowed at progression1.
The MTB integrated genomic data, clinical status, and actionability frameworks (including ESMO ESCAT) to recommend treatments, using defined variant allele frequency thresholds for tissue and liquid samples. This design allowed a unique assessment of how concordance or discordance between tissue and liquid biopsy results influences real‑world decisions and outcomes in a pan‑tumor setting1.
Concordance: how often tissue and liquid agree
In ROME, concordance was defined as the same actionable alteration being detected in both tissue and liquid biopsies and forming the basis for MTB‑recommended treatment. Under this definition, 49.2% of MTB‑indicated alterations were concordant (T+L group), while 34.8% were actionable only in tissue (T group) and 16.0% only in liquid (L group). Discordance arose from biological and technical factors, including molecular alteration discrepancies (particularly in PI3K/PTEN/AKT/mTOR and ERBB2 pathways), test failures (about 20% of discordant cases), and challenges in liquid-based biomarker detection such as TMB1.
What concordance means for patient outcomes
The most striking result from ROME is that patients with concordant actionable alterations in both tissue and liquid biopsies derived the greatest benefit from tailored therapies. In the concordant T+L group, patients receiving TT achieved1:
When focusing only on patients in the TT arm, survival followed a clear gradient1:
Patients with truly discordant molecular results had shorter overall and progression‑free survival than those with concordant profiles, supporting concordance as a potential predictive biomarker for the efficacy of matched therapy. Importantly, the survival advantage of concordant profiling was consistent across subgroups defined by tumor fraction and metastatic burden, indicating that its predictive value is robust to differences in shedding and disease extent1.
Implications for precision oncology
Findings from the ROME trial reinforce that tissue and liquid biopsies are not interchangeable but synergistic components of a modern precision oncology strategy.
Tissue biopsy remains indispensable for diagnosis, histopathology, and certain genomic events, while liquid biopsy broadens coverage across metastatic sites and over time, particularly when repeated sampling is needed. Integrating both modalities, supported by expert MTBs and comprehensive genomic profiling, maximizes the chance of detecting clinically actionable alterations and selecting patients who can truly benefit from targeted or immunotherapies1. Consistent with this, a study of 146 lung cancer patients undergoing matched tissue and plasma NGS using the same panel content reported patient‑level concordance of 83.6% and high sensitivity (≥85%) for several actionable drivers, including EGFR 19del, ALK and RET fusions, KRAS p.G12C and BRAF V600E, while also revealing subsets with tissue‑ or plasma‑specific variants4. These data underscore that liquid biopsy can function as a complementary tool to tissue biopsy, revealing both concordant variants and biopsy-specific variants when panel content is shared.
For clinical practice and trial design, ROME suggests several priorities: routinely pairing tissue and liquid profiling where feasible, explicitly considering concordance as a stratification or enrichment factor, and investing in advanced bioinformatic tools that reduce discordance and test failures1. As liquid biopsy platforms evolve and multimodal data integration matures, leveraging concordant insights from both biospecimens will be central to delivering on the promise of precision oncology for patients with advanced solid tumors.
This promise is increasingly recognized amongst healthcare experts. In a recent Delphi consensus endorsed by Italian scientific societies, most of the experts stated the importance of having both the data from tissue and liquid biopsy to integrate information from the two approaches (within a recommended ≤2-week sampling window), potentially yielding the maximum diagnostic accuracy and offering insights into the spatial-temporal heterogeneity of the disease5. However, the best testing approach – whether complementary or sequential – should be evaluated on a case-by-case basis with consideration of the patient's clinical history.
Capture the full molecular picture with SOPHiA DDM™
The SOPHiA DDM™ Platform provides seamless, end-to-end solutions for tissue and liquid biopsy, integrating powerful analytics with intuitive interpretation support to drive deeper insights and advance oncology research.
Healthcare institutions can benefit from:
With the SOPHiA DDM™ Platform, users gain access to the SOPHiA GENETICS Community, one of the largest global networks of connected healthcare institutions. Aggregated variant insights from anonymized data build collective intelligence across multiple disorders, further supporting confident, informed decision-making.
SOPHiA GENETICS products are for research use only, not for use in diagnostic procedures unless otherwise stated.
References
Discover how Centre Léon Bérard (CLB) and SOPHiA GENETICS co-developed a 21-gene liquid biopsy solution for multi-cancer analysis, designed to accurately detect both guideline-recommended and emerging biomarkers. Explore the full story to learn how this targeted approach has streamlined CLB’s analysis workflows, enabling the sequencing of 600 cell-free DNA samples per year with significantly faster turnaround times compared to traditional tissue-based methods.
In the era of precision oncology, it has become increasingly common for patients diagnosed with cancer to undergo tumor sequencing. Identifying the mutations that make up a tumor’s genomic landscape can help guide selection of targeted therapies and inform prognosis. Despite the recognized value of tumor-only sequencing, labs performing this type of testing face a number of technical challenges that, if not properly addressed, can render the results uninformative or even misleading.
Although there are a variety of inherent challenges in tumor-only sequencing, all ultimately impact the ability to accurately distinguish somatic mutations driving tumorigenesis from germline variants associated with cancer predisposition. In fact, it has been estimated that as many as one third of mutations identified by tumor-only sequencing may be false-positive germline changes, including in potentially actionable genes1. Having an accurate picture of a tumor’s genomic makeup and contextual genetic environment is crucial to an accurate clinical assessment, which impacts therapeutic recommendations and represents the patient’s best chance for successful treatment.
In this blog we explore different strategies for enriching tumor analysis for somatic mutations and discuss why matched tumor-normal sequencing has become the preferred method.
One approach is to use variants present in large population databases as a filter to remove likely germline variants from a tumor sample2. While this practice is generally effective, it will also remove true somatic variants that happen to be identical to germline variants, resulting in a false negative. Database-driven approaches can also overlook any rare germline variants missing from large population databases due to underrepresentation of non-White individuals. These variants will remain in the sequencing data and can result in false-positive germline findings.
Taking allele frequency into consideration can help. This strategy is based on the premise that an allele frequency of 50% is consistent with a heterozygous germline variant, and an allele frequency of ~100% is consistent with a homozygous germline variant1. It then stands to reason that focusing on variants with a lower allele frequency increases the likelihood of somatic origin.
While this is true, such an approach can be complicated by many factors including contamination of the tumor sample with normal tissue, tumor heterogeneity, sequencing artifacts, difficulty mapping reads in regions of high homology, high level mosaic variants that arose early in differentiation, as well as changes in allele fraction due to copy number changes or loss of heterozygosity (LOH). Any of which can lead to inaccurate attribution of origin.
Matched tumor-normal sequencing that pairs analysis of a tumor sample with that of a comparable, normal sample – most often from the same individual – has been shown to be a more effective strategy, yielding more reliable identification of the somatic changes specific to a tumor1,3,4. As the name suggests, variants in the matched normal sample are determined to be germline in origin, or of alternate origin unrelated to the current tumorigenicity. When used as a filter against the tumor sample, somatic variants relevant to the cancer at hand can be identified with a high degree of confidence. Variants found at low frequencies in the normal sample can be confidently classified as false positives if they are not significantly enriched in the tumor.
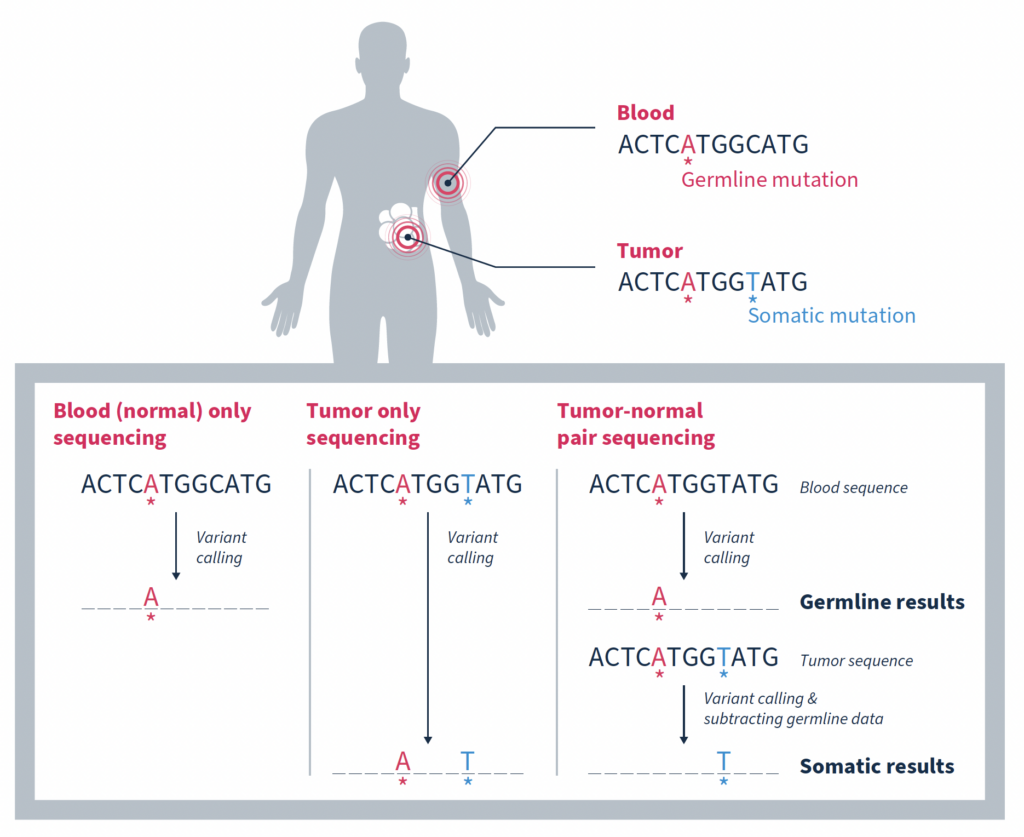
While the most important function of matched tumor-normal sequencing is to identify and retain somatic mutations, it also serves other important functions.
At the most simplistic level, biological samples can exhibit variability due to factors such as environmental influences, biological processes and sample handling. Matched-tumor normal sequencing provides a built-in baseline of background noise resulting from these factors, or from introduction of sequencing artifacts, that can be filtered out.
In the case of FFPE samples, extracted DNA is often fragmented and of a lower quality than fresh tissue samples. Matched tumor-normal sequencing provides a comparison that helps distinguish true alterations from noise resulting from degradation of the DNA, enhancing sensitivity.
Cell-free DNA (cfDNA) samples, also known as liquid biopsy samples, contain DNA from tumor cells, but they also contain a significant amount of DNA from white blood cells. In many individuals, especially those who are older, these phenotypically normal blood cells contain acquired mutations subsequently increased in relative frequency due to clonal expansion. These clonal hematopoiesis of indeterminate potential (CHIP) variants often, but not always, occur in the same genes associated with blood cancers like leukemia. However, while they are indicative of an increased risk of developing a blood cancer in the future, they are not likely to be relevant to the tumor being analyzed.
Simultaneously sequencing matched white blood cells as a normal control can successfully distinguish somatic mutations that are relevant to driving tumorigenesis from somatic mutations arising from the normal process of clonal hematopoiesis4. This is such an important consideration that both ESMO and AMP guidelines specify that matched white blood cell sequencing should be used for interpretation of somatic variants in cfDNA testing5,6.
Removal of false positives arising from CHIP variants is not only important for accurate cfDNA analysis, but also FFPE analysis. In a study by Memorial Sloan Kettering Cancer Center (MSK) investigators, matched tumor-normal sequencing results showed that 5.2% (912/17,469) of patients with advanced cancer would have had at least 1 clonal hematopoietic (CH)-associated mutation erroneously called as tumor-derived in the absence of matched blood sequencing7. Of these CH variants, 49.7% of them were classified as oncogenic or likely oncogenic based on OncoKB™, and 3.2% were associated with approved or investigational therapies (e.g. mutations in IDH1/2). Failure to recognize such mutations as blood-derived may result in inaccurate precision therapy recommendations.
The ability to distinguish between somatic and germline variants has the additional benefit of streamlining analysis of germline variants which have additional implication for a patient’s clinical care. Notably providing information about future disease risk which can be managed in part through surveillance as well as allowing for testing of family members who may also be at risk for disease.
It is for the reasons discussed here that MSK-ACCESS® powered with SOPHiA DDM™ for liquid biopsy and MSK-IMPACT® powered with SOPHiA DDM™ for comprehensive genomic profiling (CGP) utilize the matched tumor-normal analysis strategy to accurately delineate somatic variants from germline and CHIP variants.
Contact us to learn more about adopting advanced liquid biopsy and CGP technology in your laboratory.
References
What is liquid biopsy?
Liquid biopsies enable analysis of biofluids, typically blood, to examine biomarkers shed by solid tumors into circulation1. They can detect actionable genomic alterations in a non-invasive way, providing valuable insights to facilitate early cancer detection and disease monitoring2.
Tumor-derived biomarkers that are a source for liquid biopsy analysis include (Fig. 1):
CTCs are initially released from primary tumors in the tissue, travel through the bloodstream, and account for the development of metastatic tumors at distant sites in the body. As they are live cells, they have the potential to be used for functional analysis such as therapy sensitivity/resistance evaluation3. However, CTCs are rare events in the blood, which makes them difficult to identify and characterize in routine clinical practice1.
EVs are membrane-enclosed structures containing proteins, genetic material, and lipids that can provide biological information on the cell of origin4. Due to their role in pathological processes, EVs are an attractive analyte for liquid biopsy, but their isolation and purification is technically challenging1.
cfDNA refers to DNA fragments that are freely circulating in the bloodstream, primarily originating from normal cells5. Circulating tumor DNA (ctDNA) is the small portion of cfDNAthat derives from tumor cells or CTCs undergoing cell death (i.e. apoptosis or necrosis)5. There are well-established methods for isolating cfDNA, and for analyzing it using methods such as PCR and next-generation sequencing (NGS)-based tests6, making it an ideal and feasible substrate for routine genomic analysis.
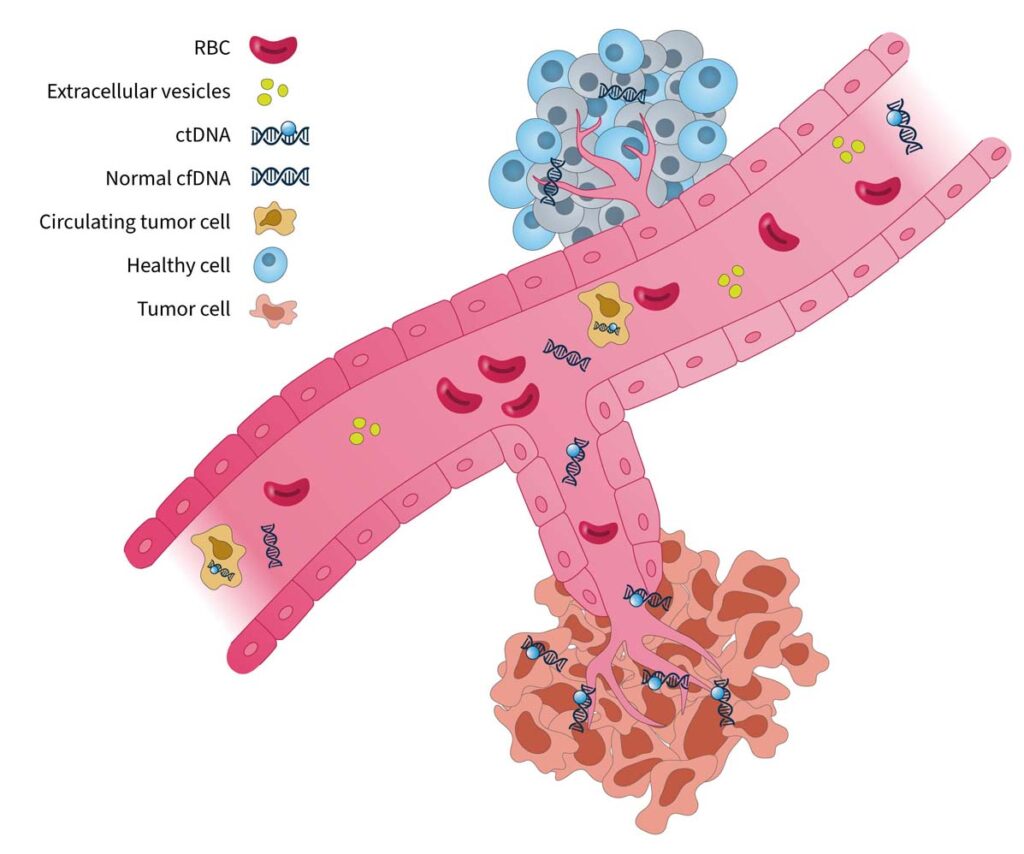
Figure 1. Blood-based cancer biomarkers in liquid biopsy7. RBC, red blood cell.
The analysis of cell-free DNA is a promising method for guiding clinical decisions and can complement current standard-of-care practices8.
What are the clinical applications of liquid biopsy?
In the era of precision medicine, tumor molecular profiling is a critical tool to identify targetable alterations and guide treatment decision-making9. Tissue biopsy is currently the gold standard for tumor profiling8; however, there are limitations associated with this approach:
Liquid biopsy has the potential to be a transformative tool in clinical oncology, showing promise for applications in many stages of cancer management (Fig. 2):

Figure 2. The advantages and clinical utility of liquid biopsy in the cancer care journey10–15.
Innovations in liquid biopsy analysis over the past decade have led to the regulatory approvals of blood-based tests to guide treatment for NSCLC, prostate, breast, and ovarian cancers16. Clinical guidelines have also provided expert recommendations for its use in specific clinical scenarios8,15. Despite great advances in technology and its increasing utility in clinical practice, there are still challenges to overcome when using liquid biopsy to identify clinically relevant information.
Overcoming “fisherman’s luck” in liquid biopsy
One challenging aspect of liquid biopsy analysis is that ctDNA concentration varies greatly across cancer types and between patients17. In patients with cancer, the quantity of ctDNA in the blood can be impacted by several factors, including histology, tumor site, clinical factors (age, sex, treatment history, etc.), and ctDNA fragmentation17. Therefore, it is important to have a robust test to detect clinically relevant variants, even at low ctDNA concentrations against a cfDNA background.
Another factor that may impact liquid biopsy analysis is the presence of clonal hematopoiesis of indeterminate potential (CHIP). In healthy individuals, the majority of cfDNA arises from hematopoietic cells (i.e. stem cells in the bone marrow that give rise to other blood cells)18. Normal hematopoietic cells accumulate somatic mutations during aging, known as CHIP, which are technically indistinguishable to tumor-specific mutations in NGS assays18,19. It is important that the biological noise caused by CHIP signals are removed in liquid biopsy analysis to eliminate false positive variant calls and give an accurate representation of disease burden19,20.
These biological confounders can make “fishing” for clinically relevant information in cfDNA a challenge (Fig. 3). For example, if a patient has high disease burden, there is likely more ctDNA available to analyze, which makes it easier to “catch” what you are looking for. However, if there is less ctDNA and more biological noise, you may need to modify your tools and approach to improve your yield.
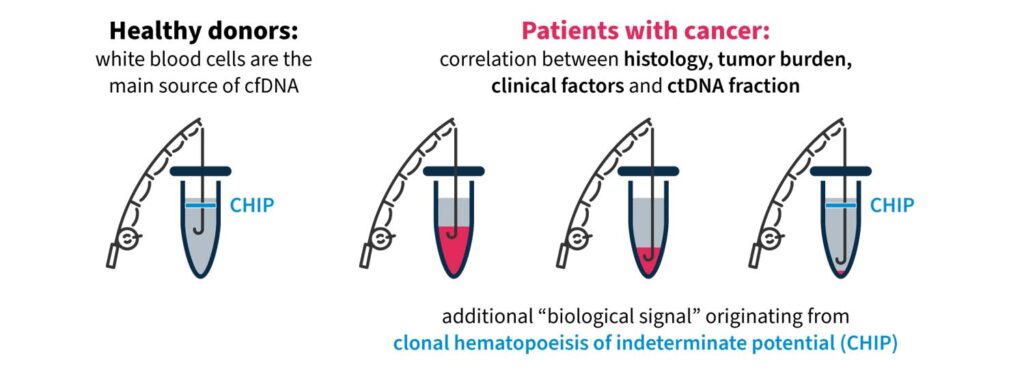
Figure 3. “Fishing” for clinically relevant information in liquid biopsy can be complicated by biological confounders17,18.
Highly precise and sensitive liquid biopsy technologies are needed to overcome “fisherman’s luck” and detect rare, causative variants and disease burden in cfDNA. Guidelines issued by the ESMO Precision Medicine Working Group on the use of cfDNA assays in clinical practice discuss the need for advanced techniques capable of capturing spatial and temporal tumor heterogeneity and reducing rates of false negatives8.
Pioneer innovation with SOPHiA DDM™ for Liquid Biopsy
SOPHiA GENETICS is at the forefront of innovation in liquid biopsy technology for tumor profiling. The advanced proprietary algorithms of the SOPHiA DDMTM Platform empower clinical researchers to reveal deep genomic insights from cell-free DNA samples.
With a streamlined, sample-to-report NGS workflow, you can:
In addition, we are excited to be collaborating with Memorial Sloan Kettering Cancer Center (MSK) to decentralize MSK-ACCESS® for liquid biopsy, designed to provide a maximum coverage of cancer disease variants in ctDNA20. By combining MSK’s clinical expertise in cancer genomics, the predictive algorithms of the SOPHiA DDM™ Platform, and the power of the global SOPHiA GENETICS community, the collaboration aims to expand access to precision cancer analysis capabilities worldwide.
Read more about how you can enhance your analytical capabilities and advance your clinical research here.
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™ Dx Hereditary Cancer Solution, SOPHiA DDM™ Dx RNAtarget Oncology Solution and SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution are available as CE-IVD products for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA DDM™ Dx Myeloid Solution and SOPHiA DDM™ Dx Solid Tumor Solution are available as CE-IVD products for In Vitro Diagnostic Use in the EEA, the United Kingdom, Switzerland, and Israel. Information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us to obtain the appropriate product information for your country of residence.
All third-party trademarks listed by SOPHiA GENETICS remain the property of their respective owners. Unless specifically identified as such, SOPHiA GENETICS’ use of third-party trademarks does not indicate any relationship, sponsorship, or endorsement between SOPHiA GENETICS and the owners of these trademarks. Any references by SOPHiA GENETICS to third-party trademarks is to identify the corresponding third-party goods and/or services and shall be considered nominative fair use under the trademark law.