Menu
Menu


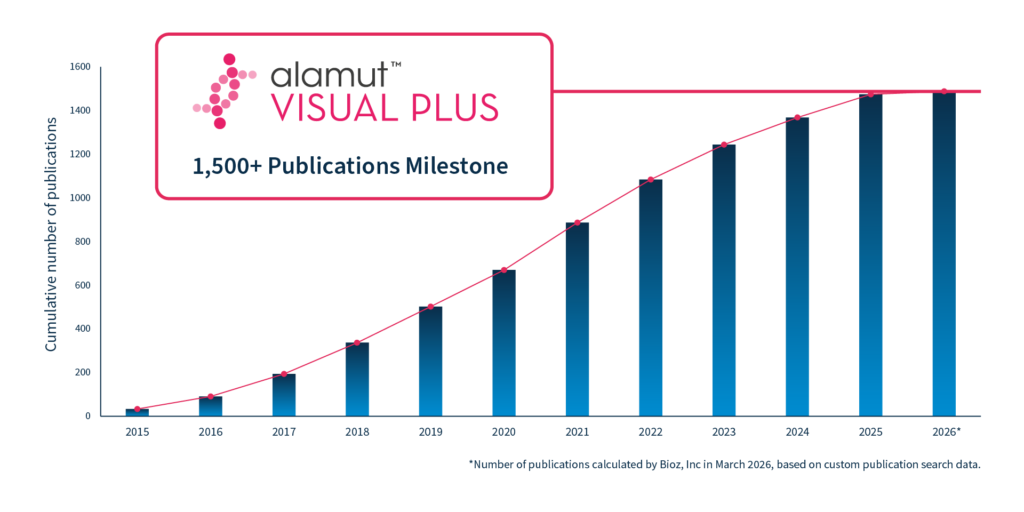
In genomics, trust in a tool is built through repeated use, transparency, and peer review. One way this trust becomes visible is through the scientific literature. Today, 1,500 peer-reviewed publications cite Alamut™ Visual Plus, marking a significant milestone in its long-standing use by the global genomics community.
These publications span a wide range of research applications, from oncology-focused analyses to investigations of rare and inherited disorders. Rather than representing a single point in time, this body of literature offers insight into how and why researchers continue to use Alamut™ Visual Plus as part of variant interpretation workflows following next-generation sequencing (NGS).
Sustained scientific relevance over time
Citations of Alamut™ Visual Plus extend across many years of genomics research, reflecting more than short-term adoption. This sustained presence suggests that researchers continue to find the software sufficiently robust, transparent, and practical to incorporate into evolving analytical workflows.
Publications citing Alamut™ Visual Plus appear in a range of high-impact, peer-reviewed journals, including:
Citation within these journals reflects the scientific rigor and scrutiny applied to the research in which the software is referenced.
As variant databases expand and interpretation frameworks mature, the continued citation of Alamut™ Visual Plus indicates ongoing relevance within the research ecosystem. Taken together, this body of literature represents long-term, high-quality engagement by the genomics community, rather than isolated or transient use.
A clearly defined role in variant interpretation workflows
A review of how Alamut™ Visual Plus is cited across the literature reveals a consistent pattern: it is most often used as a specialist variant interpretation platform, applied when deeper analysis is required beyond automated pipelines or primary annotation databases.
In particular, authors reference Alamut™ Visual Plus when addressing uncertainty around:
By aggregating multiple in silico prediction tools in one platform and contextualizing within biologically and transcript-aware frameworks, Alamut™ Visual Plus supports researchers in forming defensible functional hypotheses and interpretation rationales.
Broad use across disease areas
Publications citing Alamut™ Visual Plus span multiple areas of genomics research, reflecting the diversity of study types and disease contexts in which it is applied. Rather than being confined to a single specialty, the literature shows its use across both germline and somatic research settings.

Rare and inherited disorder applications
In studies of rare and inherited conditions, Alamut™ Visual Plus is cited in research focused on variant investigation following NGS analysis. These include studies exploring neurodevelopmental disorders, skeletal dysplasias, and inherited metabolic or syndromic conditions, where researchers must assess a wide range of candidate variants.
In this context, Alamut™ Visual Plus supports detailed examination of variants identified through variant calling pipelines.

Oncology applications
In oncology-focused genomics research, Alamut™ Visual Plus is commonly referenced in studies examining somatic variation across both solid tumors and hematological malignancies. These include research into lung, breast, endocrine, and sarcoma cancers, as well as blood-based malignancies.
Here, it appears as part of broader analytical approaches used to explore molecular characteristics of tumors, rather than as a standalone analysis tool.
Global adoption across the genomics community
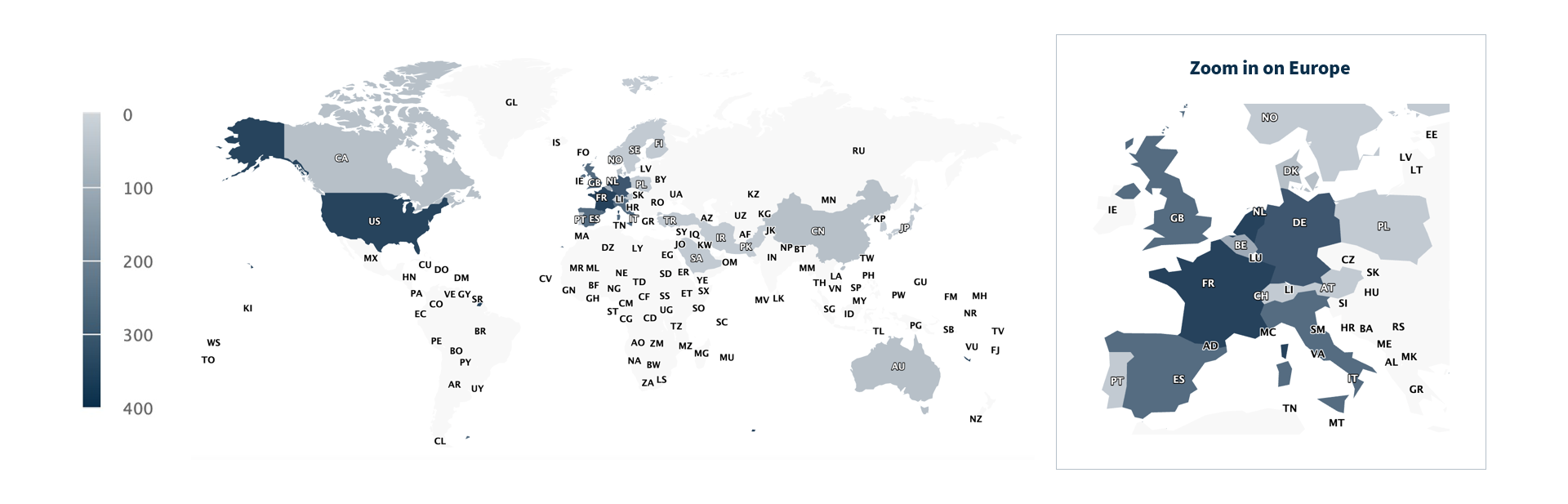
Use of Alamut™ Visual Plus is not limited to specific institutions or regions. Publications citing the software originate from groups worldwide, with notable concentrations in the United States, France, the Netherlands, Germany, Spain, the United Kingdom, and Italy.
This geographic spread underscores its role as a shared resource within the international genomics community.
Why researchers rely on Alamut™ Visual Plus in complex cases
Beyond the disease areas represented in the literature, publications also provide insight into why researchers turn to Alamut™ Visual Plus when genomic interpretation becomes challenging.
Across studies, it is most often cited when authors need to:
Rather than replacing upstream pipelines or databases, Alamut™ Visual Plus is referenced as a decision-support resource, used alongside other tools when deeper analysis or justification is required.
Key challenges addressed in genomics workflows
In silico variant effect exploration
Researchers frequently cite Alamut™ Visual Plus when investigating predicted effects of variants on splicing or protein structure. By bringing together multiple in silico prediction models within transcript-aware views, it supports comparative evaluation and hypothesis.
Variant annotation and prioritization
Another common application is the prioritization of candidate variants after initial filtering. This is particularly evident in studies addressing novel variants or variants of uncertain significance (VUS), where additional context is needed to guide further investigation or downstream functional studies.
Evolving alongside genomic advancements
The body of literature citing Alamut™ Visual Plus spans a period of rapid evolution in genomics research, including:
Continued citation across this changing landscape reflects the software’s ability to evolve alongside research practices, supported largely by active user feedback and ongoing development.
A milestone shaped by the genomics community
Each of the 1,500 publications citing Alamut™ Visual Plus represents the work of researchers applying genomic tools to investigate biological questions and disease mechanisms. This milestone therefore reflects not only the software itself, but also the global genomics community that continues to rely on it.
At SOPHiA GENETICS, supporting experts with tools designed for healthcare data exploration and variant interpretation remains a core focus.
As genomics research continues to expand in scale and complexity, tools that support careful, evidence-based interpretation remain essential. The growing number of publications citing Alamut™ Visual Plus reflects its ongoing utility across disciplines, study types, and research questions.
To learn more about the capabilities of Alamut™ Visual Plus and how it could help you to understand novel and complex variants and resolve VUS, read the tech note or visit the webpage.
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™ Dx Hereditary Cancer Solution, SOPHiA DDM™ Dx RNAtarget Oncology Solution and SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution are available as CE-IVD products for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA DDM™ Dx Myeloid Solution and SOPHiA DDM™ Dx Solid Tumor Solution are available as CE-IVD products for In Vitro Diagnostic Use in the EEA, the United Kingdom, Switzerland, and Israel. Information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us to obtain the appropriate product information for your country of residence.
All third-party trademarks listed by SOPHiA GENETICS remain the property of their respective owners. Unless specifically identified as such, SOPHiA GENETICS’ use of third-party trademarks does not indicate any relationship, sponsorship, or endorsement between SOPHiA GENETICS and the owners of these trademarks. Any references by SOPHiA GENETICS to third-party trademarks is to identify the corresponding third-party goods and/or services and shall be considered nominative fair use under the trademark law.