Menu
Menu
We are glad to host Dr. Davide Martorana, Senior Molecular Geneticist at the Medical Genetic Lab of the University-Hospital of Parma in Italy, who shared with us his institute’s experience with the adoption of the New Generation SOPHiA DDMTM Platform.
-Hello, Davide, thank you for joining us today for this spotlight! Could you please briefly introduce yourself and describe your role, institution, and the type of services you offer?
-Sure. My name is Davide Martorana, and I am a Senior Molecular Geneticist at the Medical Genetic Lab – a lab at the University Hospital of Parma in Italy.
As a Biologist, I am part of the clinical interpretation team for focused panels and clinical whole exome sequencing for subjects with a suggestive phenotype; after the genetic test, we provide reports with results interpretation.
-Could you please share with us which of the SOPHiA GENETICS applications and services are you currently using?
-We use two different SOPHiA GENETICS solutions, the Nephropathy Solutions (NES) and Hereditary Cancer Solutions (HCS), analysed with SOPHiA DDMTM
-What are the biggest benefits you see in the New Generation SOPHiA DDMTM Platform in terms of new features/ capabilities and user experience?
-First of all, I want to state that we are very happy and satisfied with this evolution of the SOPHiA DDMTM Platform. The feature I prefer is the personalization of the information flow about genetic variants; in fact, I am sure every analyst has a preferred workflow when evaluating a single variant; for example, I like to see immediately the population frequency, both in our single center and for all SOPHiA GENETICS customers; then, the ACMG-AMP classification with specified criteria.
In my opinion, this fact is very important, because when you must manage a lot of variants, it is important to focus fast on a few but essential info, then if you want there is the possibility to access several other additional info; in particular, I appreciate the extensive link-outs to other databases, which are very accurate and useful.
-Thank you for your kind words. Could you please share an example where the SOPHiA DDMTM Platform streamlined your laboratory workflows and supported your clinical research efforts?
-In a specific family, after genetic counseling of a twenty-week-old pregnant woman with a clinical diagnosis of Polycystic Kidney disease 1 - never investigated at a genetic level - we were asked to analyze the woman and the fetus with the Nephropathy solution kit.
After the analysis on the SOPHiA DDMTM Platform, we were able to detect the causative mutation in PKD1 gene in the mother but not in the fetus. The analyses were performed in just three days after receiving the biological samples, which was a very fast turnaround.
After that analysis, we realized the true potential of having an NGS kit coupled with software with fast, reliable, and accurate results, and this is just one simple example.
-It is great to see how our solutions are impacting your real-life clinical research and streamlining your decision-making when it matters the most. On the implementation side, how was your experience with the Setup program for SOPHiA DDMTM in terms of easiness, time spent on validation, and time to routine?
-In our experience, the Setup program for SOPHiA DDMTM was very fast and easy. In fact, after the training with SOPHiA customer support, we spent just a working for validating the entire process and personalizing the information flow before introducing it in the routine analysis. This is very important and efficient for the users.
-And how does your experience with the SOPHiA GENETICS customer support look like?
-I am extremely satisfied with SOPHiA GENETICS customer support because they are efficient and proactive, answering all our queries and addressing our issues in a very short time and with great competence and kindness. I would like to thank you on behalf of the entire team for that.
We would like to warmly thank Dr. Davide Martorana for joining us in this customer spotlight.
Are you interested in exploring how the New Generation SOPHiA DDM Platform can revolutionize your workflows? Check out our recent blog post here!
The term « SOPHIA » used by the speaker refers to SOPHiA GENETICS and its products. The opinions expressed during this presentation are these of the speaker and may not represent the opinions of SOPHiA GENETICS. SOPHiA GENETICS does not provide support in the validation of custom products for clinical use. SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution is available as a CE-IVD product for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
More than six people die every hour in the US from a blood cancer. Solutions can’t come fast enough for those who suffer with these cancers all around the world. Fortunately, researchers studying blood diseases have experienced rapid advances in their capabilities to develop and test effective therapies with some extremely significant advancements.
Some of the most difficult limitations of molecular profiling for hematological cancer disorders include accurate detection of mutations in GC-rich gene regions and insertions or deletions in challenging genes. Data analysis on NGS DNA samples identifies complex variants to accurately identify myeloid malignancies. This validation of targeted mutations has encouraged many medical centers to order NGS testing for every acute myeloid leukemia case.
Faster, more efficient NGS analysis can drive better hematological cancer research outcomes to potentially improve care for patients with blood cancers and diagnosis of new cases.
International guidelines for hematological cancer diagnosis and treatment are continuously evolving and create the need for laboratories’ fast adaptation. Those evidence-based guidelines by physician commissions contribute to improving the clinical standard of care. The World Health Organization, European Hematology Association, European LeukemiaNet, College of American Pathologists and the American Society of Hematology call for increased use of NGS testing for initial diagnostic workup of blood cancers.
Detection of the relevant biomarkers for myeloid malignancies by NGS, per international guidelines, helps to ensure optimal clinical trial enrollment, therapy validation, dose protocols and other research benefits. A solution that can be constantly updated and inform based on those guidelines ensures that the research is always current.
The accurate assessment of biomarkers and the validity of resulting research findings depend on reliable DNA and RNA fusion panels and easily reproducible results. Data analysis and reporting in a comprehensive platform eliminates silos of valuable data and maximizes its application.
The SOPHiA DDM™ Platform enables the upload of multimodal data from any environment to one of the world’s largest networks of connected labs. Data remains the property of the healthcare institution, but pseudonymized and pooled with like data, it can propel research and ultimately treatment forward with the goal of improved patient care.
Learn more about the capabilities of SOPHiA DDM™ Platform for myeloid biomarker detection and more by contacting us today.
In the era of precision oncology, it has become increasingly common for patients diagnosed with cancer to undergo tumor sequencing. Identifying the mutations that make up a tumor’s genomic landscape can help guide selection of targeted therapies and inform prognosis. Despite the recognized value of tumor-only sequencing, labs performing this type of testing face a number of technical challenges that, if not properly addressed, can render the results uninformative or even misleading.
Although there are a variety of inherent challenges in tumor-only sequencing, all ultimately impact the ability to accurately distinguish somatic mutations driving tumorigenesis from germline variants associated with cancer predisposition. In fact, it has been estimated that as many as one third of mutations identified by tumor-only sequencing may be false-positive germline changes, including in potentially actionable genes1. Having an accurate picture of a tumor’s genomic makeup and contextual genetic environment is crucial to an accurate clinical assessment, which impacts therapeutic recommendations and represents the patient’s best chance for successful treatment.
In this blog we explore different strategies for enriching tumor analysis for somatic mutations and discuss why matched tumor-normal sequencing has become the preferred method.
One approach is to use variants present in large population databases as a filter to remove likely germline variants from a tumor sample2. While this practice is generally effective, it will also remove true somatic variants that happen to be identical to germline variants, resulting in a false negative. Database-driven approaches can also overlook any rare germline variants missing from large population databases due to underrepresentation of non-White individuals. These variants will remain in the sequencing data and can result in false-positive germline findings.
Taking allele frequency into consideration can help. This strategy is based on the premise that an allele frequency of 50% is consistent with a heterozygous germline variant, and an allele frequency of ~100% is consistent with a homozygous germline variant1. It then stands to reason that focusing on variants with a lower allele frequency increases the likelihood of somatic origin.
While this is true, such an approach can be complicated by many factors including contamination of the tumor sample with normal tissue, tumor heterogeneity, sequencing artifacts, difficulty mapping reads in regions of high homology, high level mosaic variants that arose early in differentiation, as well as changes in allele fraction due to copy number changes or loss of heterozygosity (LOH). Any of which can lead to inaccurate attribution of origin.
Matched tumor-normal sequencing that pairs analysis of a tumor sample with that of a comparable, normal sample – most often from the same individual – has been shown to be a more effective strategy, yielding more reliable identification of the somatic changes specific to a tumor1,3,4. As the name suggests, variants in the matched normal sample are determined to be germline in origin, or of alternate origin unrelated to the current tumorigenicity. When used as a filter against the tumor sample, somatic variants relevant to the cancer at hand can be identified with a high degree of confidence. Variants found at low frequencies in the normal sample can be confidently classified as false positives if they are not significantly enriched in the tumor.
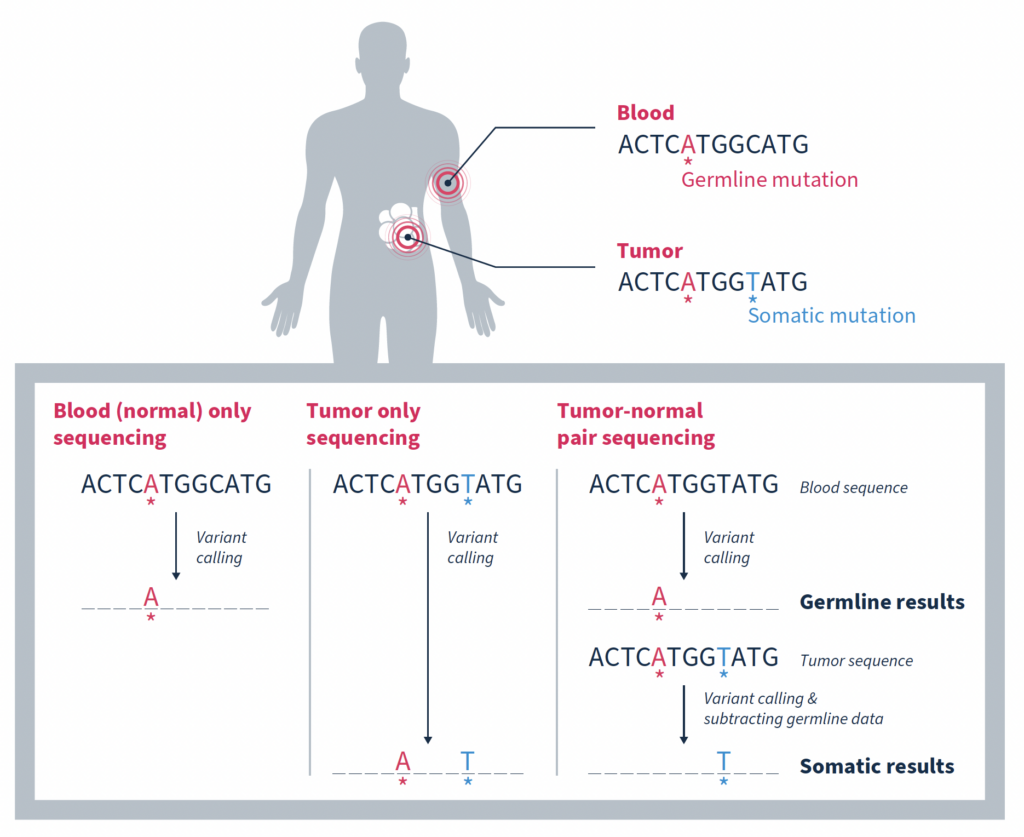
While the most important function of matched tumor-normal sequencing is to identify and retain somatic mutations, it also serves other important functions.
At the most simplistic level, biological samples can exhibit variability due to factors such as environmental influences, biological processes and sample handling. Matched-tumor normal sequencing provides a built-in baseline of background noise resulting from these factors, or from introduction of sequencing artifacts, that can be filtered out.
In the case of FFPE samples, extracted DNA is often fragmented and of a lower quality than fresh tissue samples. Matched tumor-normal sequencing provides a comparison that helps distinguish true alterations from noise resulting from degradation of the DNA, enhancing sensitivity.
Cell-free DNA (cfDNA) samples, also known as liquid biopsy samples, contain DNA from tumor cells, but they also contain a significant amount of DNA from white blood cells. In many individuals, especially those who are older, these phenotypically normal blood cells contain acquired mutations subsequently increased in relative frequency due to clonal expansion. These clonal hematopoiesis of indeterminate potential (CHIP) variants often, but not always, occur in the same genes associated with blood cancers like leukemia. However, while they are indicative of an increased risk of developing a blood cancer in the future, they are not likely to be relevant to the tumor being analyzed.
Simultaneously sequencing matched white blood cells as a normal control can successfully distinguish somatic mutations that are relevant to driving tumorigenesis from somatic mutations arising from the normal process of clonal hematopoiesis4. This is such an important consideration that both ESMO and AMP guidelines specify that matched white blood cell sequencing should be used for interpretation of somatic variants in cfDNA testing5,6.
Removal of false positives arising from CHIP variants is not only important for accurate cfDNA analysis, but also FFPE analysis. In a study by Memorial Sloan Kettering Cancer Center (MSK) investigators, matched tumor-normal sequencing results showed that 5.2% (912/17,469) of patients with advanced cancer would have had at least 1 clonal hematopoietic (CH)-associated mutation erroneously called as tumor-derived in the absence of matched blood sequencing7. Of these CH variants, 49.7% of them were classified as oncogenic or likely oncogenic based on OncoKB™, and 3.2% were associated with approved or investigational therapies (e.g. mutations in IDH1/2). Failure to recognize such mutations as blood-derived may result in inaccurate precision therapy recommendations.
The ability to distinguish between somatic and germline variants has the additional benefit of streamlining analysis of germline variants which have additional implication for a patient’s clinical care. Notably providing information about future disease risk which can be managed in part through surveillance as well as allowing for testing of family members who may also be at risk for disease.
It is for the reasons discussed here that MSK-ACCESS® powered with SOPHiA DDM™ for liquid biopsy and MSK-IMPACT® powered with SOPHiA DDM™ for comprehensive genomic profiling (CGP) utilize the matched tumor-normal analysis strategy to accurately delineate somatic variants from germline and CHIP variants.
Contact us to learn more about adopting advanced liquid biopsy and CGP technology in your laboratory.
References
Discover how Henry Ford Health streamlined their genetic testing by moving from three separate assays to a single, powerful exome-based solution. Learn how the SOPHiA DDM™ Exome Solution v3 consolidated their workflows for hereditary cancer, cystic fibrosis, and pharmacogenetics, offering broad gene coverage with a compact assay footprint. Dive into the full story to see how this innovative approach enabled detection of SNVs, Indels, and CNVs in one comprehensive assay, simplifying their germline testing processes.
Unleashing the power of healthcare data with the New Generation SOPHiA DDM™ Platform.
As the burden of cancer and rare diseases continues to grow globally, the complexity of the diseases demands more sophisticated solutions. Researchers and clinicians are constantly striving to develop novel, more effective therapies, and diagnostic tools to improve patient outcomes and resolve the biggest unmet needs in global healthcare. At the core of these efforts, there is one key element: Data.
From diagnosis to therapy selection and drug development, data is now indispensable for diagnosis and personalized treatment. The rise of precision medicine highlights the critical need for cutting-edge solutions that can harness and analyze vast amounts of healthcare data, driving advanced decision-making to improve patient outcomes at scale.
In response to this pressing need, platforms like SOPHiA DDM™ have emerged as revolutionary solutions in advancing data-driven medicine. Since its initial release in 2015, SOPHiA DDM™ has pioneered how healthcare professionals use data, having analyzed over 1.8 million genomic profiles to date and accelerating the practice of precision medicine worldwide. With nearly 30,000 analyses per month, the SOPHiA DDM™ Platform has proven itself to be a vital tool in the fight against cancer, rare and inherited diseases. Yet, as healthcare evolves, so too must the tools and technologies that support it.
In recent years, the healthcare landscape has witnessed a dramatic increase in both the volume and complexity of data. Genomic, radiomic, and clinical data have become integral to understanding diseases on a deeper level. However, the ability to process, integrate, and analyze these diverse data sources remains a significant challenge for clinicians and researchers. This challenge highlights an unmet need in global healthcare: the necessity for platforms that can break silos within and among healthcare institutions, and bridge the gap between data generation and actionable insights, allowing for more accurate diagnoses and personalized treatment strategies.
In response to this emerging need, SOPHiA GENETICS has just revealed the New Generation SOPHiA DDM™ Platform, aiming to stay at the forefront of precision medicine and address today the healthcare needs of tomorrow.
The New Generation SOPHiA DDM™ Platform not only enhances the speed and efficiency of data processing but also offers a powerful, web-based architecture designed to meet the evolving demands of clinical research. By leveraging advanced technologies like cloud computing and GPUs from world-class industry partners such as NVIDIA and Microsoft, SOPHiA DDM™ is set to revolutionize how healthcare professionals manage and interpret complex datasets to make informed decisions.
Leveraging the groundbreaking capabilities of the SOPHiA DDM™ Platform, healthcare professionals benefit from significantly reduced turnaround times, enabling quicker insights from data upload to final analysis.
Moreover, the platform’s enhanced computing capabilities allow it to process larger and more complex datasets, paving the way for new applications such as Whole Genome Sequencing (WGS), Minimal Residual Disease (MRD), Liquid Biopsy, and more, providing deeper insights into the genetic underpinnings of diseases, helping clinicians tailor therapies to individual patients with greater precision.
In addition to genomics, the platform offers advanced multimodal analytics, which are essential for understanding diseases like cancer, where multiple data types (genomic, radiomic, and clinical) need to be integrated for a more comprehensive view of the patient’s condition and unique biology. This multimodal approach allows for the analysis and interpretation of diverse data across different modalities, leading to more accurate predictions and personalized treatment plans.
One of the most significant advancements in the New Generation SOPHiA DDM™ Platform is its ability to offer genomic, radiomic, and multimodal analyses within a single, integrated workspace. This unified approach empowers healthcare providers to select the tools and applications that best suit their needs, whether they are focused on identifying genetic mutations, analyzing medical images, or integrating various data sources for predictive modeling.
By integrating these diverse data types, the New Generation SOPHiA DDM™ empowers clinicians to make better-informed decisions, improving the precision of diagnosis and treatment in oncology, rare and inherited diseases.
One of the key challenges in modern healthcare is the fragmentation of data. In many systems, vital information is siloed across different platforms and institutions, limiting the ability to generate a comprehensive understanding of a patient’s condition. SOPHiA GENETICS addresses this issue by promoting a decentralized, technology-agnostic, global platform where data can be securely shared among users, breaking down barriers to knowledge and experience exchange.
As Dr. Zhenyu Xu, Chief Scientific Officer at SOPHiA GENETICS, explains, “Our decentralized, multimodal analytics platform supports customers and helps break data silos by creating a global community where knowledge is safely and securely shared amongst users. The new generation of our SOPHiA DDM™ Platform is revolutionizing the user experience by blending our powerful AI algorithms with multimodal data to produce meaningful insights to further the field of precision medicine.”
As cancer therapies and data technologies continue to evolve, platforms like SOPHiA DDM™ will play a central role in shaping the future of precision medicine. The need for innovative, data-driven solutions is more urgent than ever, as healthcare providers strive to keep pace with the complexities of modern diseases.
Abhimanyu Verma, Chief Technology Officer at SOPHiA GENETICS, reflects on the broader impact of these advancements: “We pride ourselves on adapting our technology to meet our customers’ needs. As the technology infrastructure at most healthcare organizations worldwide has evolved, we are thrilled to continue to provide best-in-class technology and set them up for success. This new generation of our platform will allow us the flexibility to respond quickly to our customer’s evolving needs and introduce new features faster, and more efficiently.”
With its innovative architecture and advanced analytics capabilities, the new SOPHiA DDM™ Platform represents a major leap forward in precision medicine. By addressing the unmet needs in global healthcare data analysis, SOPHiA GENETICS is helping to pioneer a future where data-driven insights lead to more personalized, effective, and timely care for patients around the world.
Learn more about the New Generation SOPHiA DDM™ Platform here. Interested in getting a free demo of the Platform? Book it here!
The ESMO Precision Medicine Working Group (PMWG) first published its recommendations for when to use next-generation sequencing (NGS) in routine practice for patients with metastatic cancers in 20201. At that time, based on identification of recurrent genomic alterations in the eight most deadly cancers and their ranking on the ESMO Scale for Clinical Actionability of molecular Targets (ESCAT), NGS was recommended for advanced non-squamous non-small-cell lung cancer (NSCLC), prostate cancer, ovarian cancer and cholangiocarcinoma. It was additionally noted as an alternative to PCR for advanced colorectal cancers.
Advances in precision medicine during the past four years have resulted in revised ESCAT rankings for a number of biomarkers, leading the PMWG to reconfirm their previous recommendations and to expand the set of metastatic cancers recommended for NGS testing. The recommendations now include NGS testing for advanced breast cancer as well as the following advanced rare cancers: gastrointestinal stromal tumor, soft-tissue sarcomas, thyroid cancer2.
Due to the potential of NGS to help identify the primary tumor, plus its positive impact on patient outcomes, the PMWG additionally recommends that NGS testing be performed for cancers of unknown primary.
In countries where targeted therapies are accessible, recommendations for NGS testing now further extend to detection of the following tumor-agnostic biomarkers:
The PMWG stresses the importance of carrying out NGS testing in clinical research centers as well as ensuring that the selected test includes the actionable biomarkers of interest. They specifically flag the importance of assessing the chosen assay for its ability to detect fusions and homologous recombination deficiency (HRD), when relevant, as not all assays have these capabilities. The PMWG also highlight the assay’s ability to identify clonal hematopoiesis (CH) as an important consideration. High-risk CH can be found in patients with solid tumors, especially when plasma cell-free DNA sequencing is performed. To be considered as CH of indeterminate potential (CHIP), the somatic variants of haematological malignancy-associated genes should be with a variant allele fraction (VAF) of ≥2%.
See the full set of guidelines here: ESMO Recommendations for the use of NGS for patients with advanced cancer.
At SOPHiA GENETICS, we offer solutions that cover the major biomarkers highlighted in the ESMO recommendations, including the key fusions noted above. Moreover, the PMWG recognizes SOPHiA DDM™ Dx HRD Solution as one of only a few methods validated for HRD detection in advanced ovarian cancer.
Additionally, we offer liquid biopsy and solid tumor applications that leverage a matched tumor-normal sequencing approach to identify CHIP and germline variants, revealing genomic alterations of true somatic origin. Recent ESMO guidelines on reporting genomic test results for solid cancers recommend that, when feasible and with proper consent, the NGS report should specify whether alterations are of germline versus somatic origin3. With matched tumor-normal sequencing, the germline origin of any variant can be determined with certainty3.
Read our flyer for further information on how SOPHiA DDM™ for Solid Tumors advances clinical research by aligning with guideline recommendations: View the flyer
References
Tell us a bit about your laboratory.
The Genetics and Genomics Laboratory in which we operate belongs to a Local Health Authority in the Sardinia region. We provide molecular evaluation of gene alterations in many solid neoplasms to identify patients eligible for personalized treatments (i.e. targeted therapy). This service is provided to medical oncology departments in the city of Cagliari and many peripheral areas in the region.
What are the roots of your collaboration with SOPHiA GENETICS, and which application are you using?
The decision to collaborate with SOPHiA GENETICS was made in 2019, as our request to activate our NGS service was subject to CE-IVD certification of the library preparation and sequencing path. Furthermore, the process seemed quite linear to us.
We are currently using the SOPHiA DDM™ RNAtarget Oncology Solution (ROS) for RNA sequencing.
Can you describe your experience of using SOPHiA DDM™ ROS in routine for your RNA sequencing?
It has an excellent ability to identify fusion gene events involved in many oncological diseases. The application allowed us to identify several known and unknown molecular alterations (single nucleotide variants, Indels, gene fusion and exon skipping events) involved in various cancers, which represent biomarkers for approved and agnostic target drugs. In this way, it is possible to perform a combined analysis of different tumor pathologies in a single analysis session.
“It has an excellent ability to identify fusion gene events involved in many oncological diseases.”
Can you describe your experience with automating the workflow?
The automation of the workflow for library preparation has allowed us to standardize the quality (e.g. coverage of control genes, average size of transcripts, coverage of target regions).
In addition, we have solved the problem of complexity in library preparation. The three days of work required for preparation of libraries, up to the loading of the pools, have been reduced to only a few steps. The manual procedure now involves the set-up of the automation tool, loading of the pools into the sequencer, and approx. 2–3 days for the analysis of the data (for 16–24 samples).
“[With automation], we have solved the problem of complexity in library preparation. The three days of work required for preparation of libraries have been reduced to only a few steps.”
What are the greatest benefits of using SOPHiA DDM™ ROS?
The greatest benefit of this application is the advanced variant filtering capability and pathogenicity pre-classification that optimizes the data analysis flow and facilitates data interpretation.
In addition, the creation of a particularly exhaustive report for clinical information and ongoing trials relating to the pathologies under examination, make the results usable by the clinician for the therapeutic choice and prognosis of the disease.
We’d like to thank Manuela Badiali, Rita Congiu, and Stefania Murru for their time and for sharing their experience. Click here to learn more about SOPHiA DDM™ RNAtarget Technology and request a demo!
The Laboratory of Genetics and Genomics use a CE-IVD version of SOPHiA DDM™ ROS called SOPHiA DDM™ Dx ROS, available in in the European Economic Area (EEA), the United Kingdom and Switzerland. The CE-IVD application is not designed for use with automation – The Laboratory of Genetics and Genomics have validated the automated workflow for clinical use. SOPHiA GENETICS products are for Research Use Only, not for use in diagnostic procedures unless otherwise specified.
We were delighted to meet with the two presenters of this poster, Elizabeth Ratsma, Pre-Registration Clinical Scientist - Cancer Genomics and Charlotte Flanagan, PhD, Innovation Lead at The Royal Marsden NHS Foundation Trust. The poster was recently presented at the ESMO Gynaecological Cancers Congress 2024 in Florence, Italy.
We would like to warmly thank them for their insightful presentation and for sharing the key takeaways of their study and experience with us through this following interview.
Dear Elizabeth and Charlotte, could you please share with us more information about the scope of this study? What was the aim of it?
Homologous Recombination Deficiency (HRD) testing is available to patients in the NHS who have been newly diagnosed with high-grade epithelial ovarian cancer.
Prior to December 1, 2023, eligible patient samples were sent to the United States for testing via Myriad. The aim of this study was for the North Thames Genomic Laboratory Hub to establish an alternative in-house method for testing ovarian FFPE samples for genomic instability and BRCA mutation status. This would provide an HRD score to enable patients to access PARP inhibitors.
How was your experience with implementing the SOPHiA DDM™ GIInger Genomic Integrity Solution?
We had experience using the SOPHiA DDM™ Platform from previous demos trialing the SOPHiA DDM™ Dx HRD Solution. The SOPHiA DDM™ GIInger Genomic Integrity Solution (or GIInger™) is accessed in a similar process, and it was not difficult to navigate this new workflow in the software. Our bioinformatics team developed a CLC script with support from SOPHiA GENETICS to upload our low-copy WGS sample FASTQ files into the SOPHiA DDM™ Platform, where they are analyzed and genomic instability scores are generated.
SOPHiA GENETICS were receptive to our suggestions to help optimize functionality for use in a clinical diagnostic setting. For example, updating the format and accessibility of reports to facilitate pairing the results with our in-house tBRCA reporting (RMH200, Roche analyzed using DRAGEN, Illumina) and LIMs system.
We have received good technical support when needed through the SOPHiA GENETICS JIRA system.
"We are looking forward to future updates, including a web-based portal and automated download of result files for our clinical scientists to access with ease."
In the scope of this study, you compared GIInger™ and the SOPHiA DDM™ Dx HRD Solution. Could you please summarize the purpose and outcomes of this comparison?
The SOPHiA DDM™ Dx HRD Solution is a CE-marked HRD solution that utilizes SOPHiA GENETICS’ preferred chemistry for whole genome and capture library preparations. At Royal Marsden Hospital (RMH), we had previously implemented a robust NGS protocol (RMH200) capable of producing BRCA capture and low-copy whole genome libraries. To optimize operational efficiency, we decided to explore using our existing automated chemistry and library preparation workflow and opt for a bioinformatics solution to analyze the whole genome data for genomic instability to provide a HRD status.
To this end, 23 samples (previously tested via an orthogonal method) were sequenced using our in-house NGS chemistry and analyzed using GIInger™ paired with our in-house tBRCA calling and compared to the SOPHiA DDM™ Dx HRD Solution, which utilizes different chemistry and a full bioinformatics solution. Reassuringly, we found 100% concordance.
Thank you for sharing these insights with us! To conclude this spotlight, we would like you to share the key takeaways of this study.
Utilizing the SOPHiA GENETICS GIInger™ bioinformatics solution paired with our in-house RMH200 panel for tBRCA status, RMH successfully launched in-house HRD testing in December 2023. By April 1, 2024, we had tested 106 samples internally, achieving recent average turnaround times of less than 21 days.
"The support provided by SOPHiA GENETICS has been sufficient and rapid, which has been invaluable during the first six months of this new service."
We would like to thank Elisabeth and Charlotte for their participation in this spotlight.
Learn more about:
SOPHiA DDM™ GIInger Genomic Integrity Solution
SOPHiA DDM™ Dx Homologous Recombination Deficiency (HRD) Solution
Disclaimer
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise. The SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution is available as CE-IVD product for In Vitro Diagnostic Use in European Economic Area (EEA), the United Kingdom and Switzerland.
The transition from the In Vitro Diagnostic Directive (IVDD) to the In Vitro Diagnostic Regulation (IVDR) in the European Union marks an important advancement in regulatory standards for genetic testing and analysis. The new standards promote transparency and traceability throughout genomic analysis processes, helping to ensure the reliability and accuracy of diagnostic results and ultimately patient safety.
Everyone wants to ensure that genomic analysis is safe for patients. However, healthcare institutions face real challenges in transitioning from IVDD to IVDR. Particularly with the use of complex software solutions for genomic data analysis, many of which are designated as research use only (RUO).
One of the biggest changes for healthcare institutions is that IVDR specifically regulates in-house manufactured tests. With few exceptions, healthcare institutions utilizing in-house manufactured tests must now meet the same requirements and proof of conformity with IVDR as manufacturers. These requirements extend to the software that is used for the analysis, interpretation and reporting of NGS data. Software developed in-house, including from public domain materials, must meet many of the same requirements and proof of conformity as commercial software solutions.
Here we answer some of the most pressing questions about what IVDR compliance entails for healthcare institutions performing genomic analysis and how CE-IVD certified software solutions can help.
Q: How does IVDR impact the analysis, interpretation, and reporting of NGS data?
A: Software used to support the analysis, interpretation and reporting of NGS data from genetic testing must conform to IVDR’s general safety and performance requirements (GSPR) to ensure reliability and safety.
Q: What are the key requirements for meeting IVDR compliance?
A: Genetic tests and their analytical software require that a healthcare institution perform the following to ensure IVDR compliance:
It is also important to ensure and demonstrate that suppliers are complying with applicable regulatory requirements.
After launch, post-market surveillance is required to monitor the safety and clinical performance of the test. Genetic tests require regulatory management of updates as well as reporting of any serious incidents, with the corrective actions taken.
Q: When does IVDR take effect?
A: IVDR replaced IVDD in 2022, with timelines for compliance differing for devices with new, versus legacy, status and class. Genetic tests and analytical software that are Class C devices, for example, must comply with IVDR by 2028, and in-house manufactured devices must comply with all relevant IVDR requirements by 2030 at the latest (see timeline).
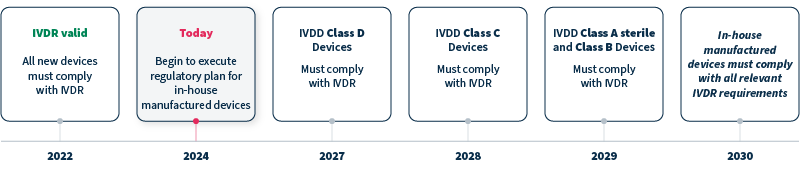
Q: How can commercial software solutions help with the IVDR transition?
A: When using a CE-IVD commercial software solution, the manufacturer’s CE-IVD certification and their existing technical documentation provide proof of compliance with current regulations, helping to lessen the burden for the healthcare institute.
Q: Can research use only (RUO) solutions be used to meet IVDR requirements?
A: RUO solutions are not intended or validated for clinical diagnostic use. RUO solutions will not be considered compliant without additional testing and validation as part of an in-house manufactured device.
Q: What are the most important considerations when evaluating commercial CE-IVD analytical software solutions?
A: When evaluating analytical software solutions, regulatory compliance is a must. However, it is also important to consider analytical and clinical validity. Does the solution provide reliable data processing and variant calling with high sensitivity and specificity? What variant types does it cover? Is it able to interpret variants according to established guidelines? Looking ahead to the future, can the solution scale with growing genomic analysis volumes?
As important as the new IVDR standards are to ensuring patient safety, it is clear that the transition poses a number of challenges to healthcare institutions performing genomic analysis. At SOPHiA GENETICS we’re proud to help simplify the transition, offering fast, reliable CE-IVD oncology applications powered by SOPHiA DDM™.
To learn more, explore our current portfolio and stay tuned for further updates.
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™ Dx Hereditary Cancer Solution, SOPHiA DDM™ Dx RNAtarget Oncology Solution and SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution are available as CE-IVD products for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA DDM™ Dx Myeloid Solution and SOPHiA DDM™ Dx Solid Tumor Solution are available as CE-IVD products for In Vitro Diagnostic Use in the EEA, the United Kingdom, Switzerland, and Israel. Information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us to obtain the appropriate product information for your country of residence.
All third-party trademarks listed by SOPHiA GENETICS remain the property of their respective owners. Unless specifically identified as such, SOPHiA GENETICS’ use of third-party trademarks does not indicate any relationship, sponsorship, or endorsement between SOPHiA GENETICS and the owners of these trademarks. Any references by SOPHiA GENETICS to third-party trademarks is to identify the corresponding third-party goods and/or services and shall be considered nominative fair use under the trademark law.