Menu
Menu
Precision medicine is transforming healthcare, particularly in oncology, by enabling treatment strategies tailored to the unique characteristics of each tumor. At the heart of this shift lies high-quality biomarker testing, driven by advanced technologies like next-generation sequencing (NGS). As NGS becomes increasingly essential for the successful development and commercialization of innovative therapies, it plays a key role in driving enhanced patient outcomes.
In this audio, Jess Lambe, VP & Managing Director BioPharma Business Development, SOPHiA GENETICS, and Louis Welebob, Ph.D, SVP, Companion Diagnostics, Myriad Genetics, uncover the details of our groundbreaking collaboration. Together, they explain how our unique hybrid testing approach has the potential to accelerate innovation and drive meaningful clinical and commercial impact, reshaping the future of cancer care.
In this session, you will:
Disclaimer notice
© 2026 SOPHiA GENETICS. All rights reserved.
The term SOPHiA used by the speakers refers to SOPHiA GENETICS and its products.
The opinions expressed during this presentation are those of the speakers and may not represent the opinions of SOPHiA GENETICS.
SOPHiA GENETICS does not provide support in the validation of custom products for clinical use.
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise. This session contains information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us at [email protected] to obtain the appropriate product information for your country of residence.
The rise of biomarker-driven therapies created a need for more precise diagnostics. As such, CDx have become indispensable for identifying patients likely to respond to a treatment.
Historically, pharmaceutical companies have pursued two main CDx development and regulatory pathways: the single-site pre-market approval (ssPMA) and the distributed in vitro diagnostic (IVD) kit approaches.
The ssPMA model leverages a single laboratory to develop, validate, and perform analytical and clinical validation of the diagnostic test. Because it offers speed, flexibility, and reduced upfront investment, it is particularly valuable in early drug development when therapeutic efficacy is still being established. But its commercial reach is limited since testing is confined to one laboratory, limiting patient access.
By contrast, the distributed IVD kit model is designed for scale. Diagnostic manufacturers develop and validate kits ready for deployment across multiple laboratories worldwide, enabling broad patient access at therapeutic launch. The drawback is the higher cost and longer timelines required to align full IVD development with drug approvals.
Pharmaceutical companies often treated these models as mutually exclusive, yet both have strengths that can be combined to meet today’s market needs.
The future of CDx development lies in a hybrid development and deployment model that combines centralized laboratory validation and clinical trial enrolment with decentralized testing at the point of care. By integrating ssPMA speed and IVD scalability, pharma companies can better align diagnostics with drug development timelines while ensuring broad access at launch.
By partnering with diagnostic companies, pharma can start the traditional central lab approach to develop the clinical trial assay (CTA), while the diagnostic partner simultaneously prepares the distributed IVD for decentralized testing. Through comparison or bridging studies, the ssPMA-approved assay can evolve into a globally distributed kit. This ensures alignment with drug development timelines and expands patient access from day one.
A strong example of this hybrid model is the partnership between SOPHiA GENETICS and Myriad Genetics to develop MSK-ACCESS® powered with SOPHiA DDM™ into a CDx test. Combining the strengths of each, the companies will start by developing a CTA and pursuing FDA approval, which will later transition into an IVDR-certified, globally deployed kit via bridging studies.
This unique CDx concept can be implemented across multiple platforms and laboratories, ensuring maximum patient access, including underserved regions. It also illustrates how a hybrid deployment accelerates timelines, increases scalability, and strengthens equity in precision oncology.
Precision medicine is in its infancy, but as it continues to grow, so must the strategies to support equitable access to CDx. The traditional pathways for CDx development are no longer fulfilling the scientific, clinical, and commercial needs of the industry. The future depends on collaboration between pharma and diagnostics companies to implement strategies, such as a hybrid approach to CDx, that accelerate innovation, ensure regulatory alignment, and deliver local access at global scale.
As demonstrated by SOPHiA GENETICS and Myriad Genetics, the industry is moving toward smarter, more efficient, and collaborative solutions. Harnessing the strengths of a dual, centralized and decentralized approach allows pharma to combine efficiency, speed, and equity, redefining the future of CDx and expanding the reach of precision oncology worldwide.
Just two decades ago, cancer was largely considered an organ-based disease. For example, lung cancer, despite having known histological subtypes, was uniformly treated as a single disease – chemotherapy for all. Today, advances in clinical genomics have transformed lung cancer into a collection of rare diseases defined by a long tail of distinct genomic alterations. Building on this knowledge, targeted therapies have gradually improved patient outcomes for eligible patients. However, significant medical needs remain, particularly regarding overall survival (OS) or achieving durable cures. Most lung cancer patients in first-line therapy do not have a clear molecularly driven cause and almost uniformly receive an immunotherapy-based treatment as a standard of care. Despite massive investments, single-biomarker approaches have failed to reliably predict response to immunotherapy, leaving clinicians unable to determine which patients will benefit most.
To address these challenges, precision medicine must shift from a siloed, single-biomarker approach to a more integrated multimodal approach, combining genomic, imaging, clinical, and biological data. Intuitively, taking a more holistic view of the patient, the tumor, and the host environment should open a stronger window into the biology of health and disease. Yet, realizing the full potential of this approach requires a transformation of the underlying data infrastructure. This includes breaking down silos across data modalities, standardizing and harmonizing datasets and promoting real-world knowledge sharing across institutions. Such transformation is necessary to unlock the power of artificial intelligence (AI) applied to multimodal healthcare data.
In this article, we explore the groundbreaking potential of multimodal AI-driven technology as a key driver of a new era in precision medicine. This paradigm shift promises not only to accelerate innovation but also to improve patient outcomes and expand equitable access to care. With competition intensifying and regulatory scrutiny increasing, biopharma companies must embrace this transformation to stay ahead and reshape the future of precision oncology.
Over the past decade, we have seen a dual revolution in healthcare: the explosion of multiple types of digital health data being produced at scale in clinical routine (e.g., genomics, imaging, electronic health records (EHR) entries) and generational breakthroughs in analytics capabilities (e.g., machine learning, deep learning, foundational models). In theory, this combination should have unlocked the full potential of precision medicine, but in practice, we are arguably still at its Stone Age. One of the root causes of this lies in the fact that healthcare data remains largely fragmented and unharmonized, and the tools to integrate and harness it effectively are often lacking. Additionally, there is no built-in incentive in the ecosystem for large-scale data and insights sharing across institutions, while preserving privacy. The “publish or perish” mantra in academia still tends to encourage data hoarding — consciously or not — further hindering collaboration.
Similarly, on the biopharma side, we see increasing interest and expertise for AI-driven approaches on specific data modalities (e.g., digital pathology, radiology). Yet, these initiatives are still rarely connected in a truly multimodal framework. Proprietary clinical trial databases remain challenging to harmonize and merge together due to heterogeneous patient consents, compliance issues, and required investments.
A fair question to ask is, therefore—is it even worth the trouble?
Although we are just beginning to tap into the capabilities of multimodal AI-driven technology, it is already driving significant advances in our understanding of health and disease. By seamlessly integrating and analyzing diverse data sources, this approach enables a more holistic perspective of complex diseases like cancer, and the patient beyond the disease.
The practical application of AI-powered multimodal technology can help biopharma companies solve complicated biological puzzles and overall optimize the drug development process (Figure 1).
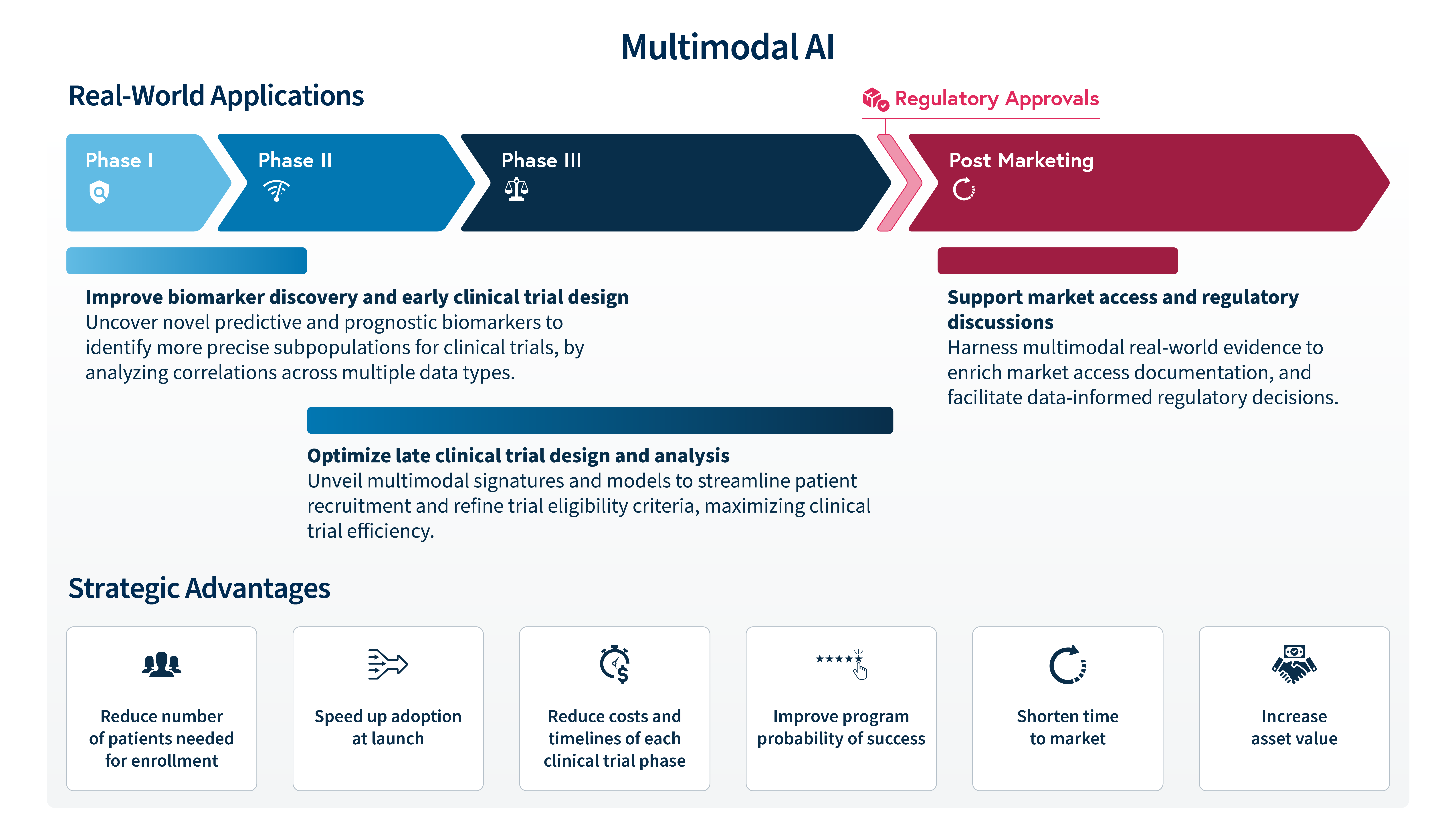
At SOPHiA GENETICS we believe that multimodal AI is no longer an option but a necessity to accelerate precision medicine. Our cloud-based SOPHiA DDM™ Platform seamlessly integrates and standardizes diverse data types into a unified analytical framework, comprising state-of-the-art specialized computational modules for data processing and analysis (e.g., genomics, radiomics), including a dedicated multimodal factory. This engine combines, extracts, and structures complex multimodal data to fuel the development of predictive analytics, delivering actionable insights that empower data-driven decision-making through an intuitive, user-friendly interface (Figure 2).

The potential of this multimodal approach is evident in the initiatives we are leading here at SOPHiA GENETICS. One noteworthy example is the TRIDENT project, presented at ESMO 2024 (Skoulidis et al., 2024). TRIDENT was a retrospective multimodal re-analysis of AstraZeneca’s Phase 3 POSEIDON trial (NCT03164616). AI-powered predictive models of treatment benefit were trained on the totality of the clinical trial data, including clinical, biological, imaging and genomics data, with the intent to identify patient subpopulations that may derive greater benefit from the addition of a CTLA-4 inhibitor on top of a PD-L1 and chemotherapy backbone in first-line non-small cell lung cancer. These models yielded signatures identifying approximately 50% of the trial population in scope that would be predicted to benefit from the addition of CTLA-4 inhibition, with a hazard ratio reduction from 0.88 (95% CI, 0.68-1.12) to 0.56 (95% CI, 0.33-0.97) in the non-squamous histology population (Figure 3). These multimodal signatures are clinically interpretable and can be readily deployed in the real-world setting on the SOPHiA DDM™ Platform for further clinical research.

This proof-of-concept analysis highlights a fundamental observation: traditional methods of analyzing and making sense of existing data, for example, through univariate or multivariate analyses, can leave significant portions of the full picture unseen. In contrast, new multimodal approaches have the potential to reveal insights that would otherwise remain hidden.
The transformational findings from this lung cancer project are not an exception. Similar results were obtained in other cancer types, such as in kidney cancer (Boulenger de Hauteclocque et al., 2023; Margue et al., 2024), and breast cancer (Groheux et al., 2025).
To successfully implement a multimodal AI-driven strategy, it is essential to begin with a clearly defined clinical question. What are you trying to predict or stratify, and why does it matter clinically? What is your endpoint of interest? These considerations will dictate the type, quality and volume of data required for a successful analysis.
A truly multimodal model demands a deep understanding of the signal-to-noise ratio within each individual data modality. For example, in genomics: do you know how the DNA of the tumor was sequenced, which chemistry was used, and on which sequencing platform? How was the variant calling performed, and what are the known limitations? Only with this level of detail can you distinguish meaningful biological signals from background noise. The same applies to radiology, where harmonizing image data across different platforms and reconstruction techniques is essential. Skipping these foundational data preparation steps risks identifying patterns in noise rather than signal.
Once the dataset is well understood and curated, data augmentation can be applied—for instance, through radiomics analysis of 3D medical imaging or pathway analysis of relevant genomic variants.
After augmentation, data aggregation becomes the next critical step. This involves integrating diverse data types into a unified analytical framework. From here, various statistical learning methods can be selected based on the clinical objective. Typically, imputation techniques for missing data need to take place before feature selection, statistical model selection, and calibration. Finally, ensuring analysis interpretability, both at the cohort and individual level is an important step in facilitating discussions with oncologists and other healthcare professionals concerning the methodology and the outcomes of the models (Figure 4). This can be achieved by using traditional machine learning models.
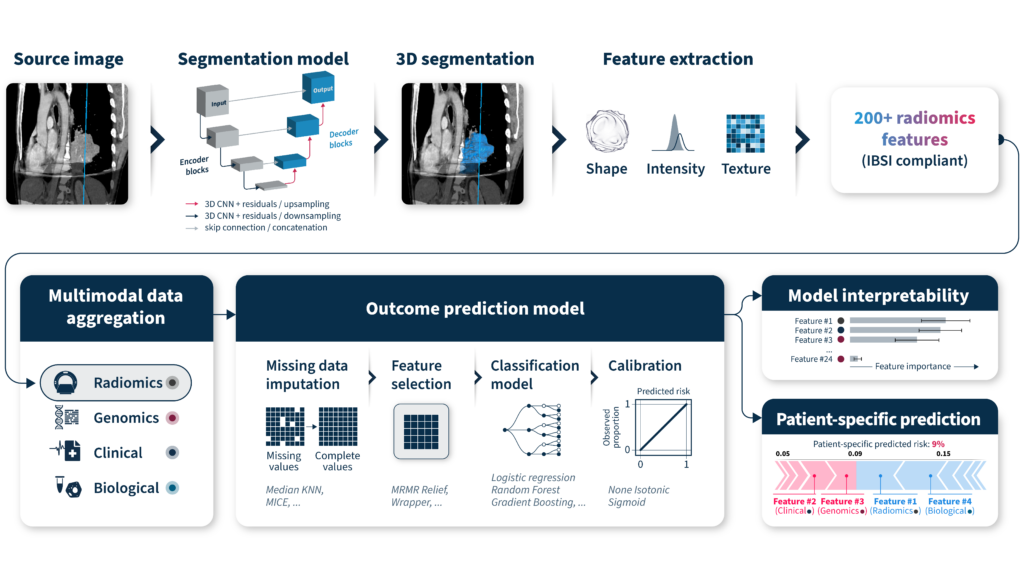
An often overlooked yet crucial step is the deployment of the models in a real-world setting. How will end users interact with the model? How will data be input and managed? What safeguards are in place for data privacy and security, and how will computational infrastructure scale?
At SOPHiA GENETICS, we believe that the cloud-native SOPHiA DDM™ Platform is uniquely positioned to spearhead this movement. Already adopted by over 800 healthcare institutions across more than 70 countries, the platform has securely processed data from more than 2 million patients, ensuring privacy while enabling impactful, AI-powered multimodal insights at scale.
The transition from single-modality to multimodal AI-driven analysis represents a paradigm shift in precision medicine. Organizations that successfully integrate diverse data modalities and multimodal technology will be best positioned to drive better patient outcomes and maximize drug development success.
Realizing this potential, however, demands more than technical innovation. It demands systemic transformation across the overall healthcare landscape — from evolving regulatory frameworks for multimodal CDx, to updated reimbursement models, standardized deployment practices— inclusive of post-market surveillance—, and greater education for both clinicians and patients.
For biopharma companies, the path forward is clear:
About twenty years ago, cancer was still considered an organ disease. Looking back today, this may look like distant, medieval times. Twenty years from now, new generations of life sciences professionals may look at 2025 in a disturbingly similar way. The multimodal revolution is only now getting started.
Written by Philippe Menu, MD, PhD, MBA - EVP, Chief Medical Officer & Chief Product Officer, SOPHiA GENETICS
References
Skoulidis F et al. 1325P TRIDENT: Machine learning (ML) multimodal signatures to identify patients that would benefit most from Tremelimumab (T) addition to durvalumab (D) + chemotherapy (CT) with data from the POSEIDON trial. Ann. Oncol. 35, S842–S843 (2024).
Boulenger de Hauteclocque A et al. Machine-learning approach for prediction of pT3a upstaging and outcomes of localizaed renal cell carcinoma (UroCCR-15). BJU Int. 2023; 132(2):160-169. doi: 10.1111/bju.15959.
Margue G et al. UroPredict: Machine learning model on real-world data for prediction of kidney cancer recurrence (UroCCR-120). NPJ Precis Oncol. 2024; 8(1):45. doi: 10.1038/s41698-024-00532-x.
Groheux D et al. FDG-PET/CT and Multimodal Machine Learning Model Prediction of Pathological Complete Response to Neoadjuvant Chemotherapy in Triple-Negative Breast Cancer. Cancers. 2025; 17(7):1249. doi: 10.3390/cancers17071249.
More than six people die every hour in the US from a blood cancer. Solutions can’t come fast enough for those who suffer with these cancers all around the world. Fortunately, researchers studying blood diseases have experienced rapid advances in their capabilities to develop and test effective therapies with some extremely significant advancements.
Some of the most difficult limitations of molecular profiling for hematological cancer disorders include accurate detection of mutations in GC-rich gene regions and insertions or deletions in challenging genes. Data analysis on NGS DNA samples identifies complex variants to accurately identify myeloid malignancies. This validation of targeted mutations has encouraged many medical centers to order NGS testing for every acute myeloid leukemia case.
Faster, more efficient NGS analysis can drive better hematological cancer research outcomes to potentially improve care for patients with blood cancers and diagnosis of new cases.
International guidelines for hematological cancer diagnosis and treatment are continuously evolving and create the need for laboratories’ fast adaptation. Those evidence-based guidelines by physician commissions contribute to improving the clinical standard of care. The World Health Organization, European Hematology Association, European LeukemiaNet, College of American Pathologists and the American Society of Hematology call for increased use of NGS testing for initial diagnostic workup of blood cancers.
Detection of the relevant biomarkers for myeloid malignancies by NGS, per international guidelines, helps to ensure optimal clinical trial enrollment, therapy validation, dose protocols and other research benefits. A solution that can be constantly updated and inform based on those guidelines ensures that the research is always current.
The accurate assessment of biomarkers and the validity of resulting research findings depend on reliable DNA and RNA fusion panels and easily reproducible results. Data analysis and reporting in a comprehensive platform eliminates silos of valuable data and maximizes its application.
The SOPHiA DDM™ Platform enables the upload of multimodal data from any environment to one of the world’s largest networks of connected labs. Data remains the property of the healthcare institution, but pseudonymized and pooled with like data, it can propel research and ultimately treatment forward with the goal of improved patient care.
Learn more about the capabilities of SOPHiA DDM™ Platform for myeloid biomarker detection and more by contacting us today.
Predictive analytics refers to using statistical algorithms, machine learning techniques, and historical data to forecast future events. In clinical trials, this means integrating diverse data sources – such as, past clinical trials, patient records, and real-world evidence (RWE) – to provide more accurate predictions about trial outcomes, patient responses, and potential risks3. By analyzing patterns from this data, predictive analytics offers a powerful tool for improving the efficiency, accuracy, and safety of clinical trials1.
Traditionally, clinical trials have been labor-intensive and costly, often taking years to yield results4. Researchers had to rely on historical outcomes, guesswork, and incomplete data to design trials and predict success. Predictive analytics changes this paradigm by enabling data-driven decision-making5. By analyzing data from clinical trials, and real-world data (RWD) – including but not limited to patient demographics, electronic health records (EHRs), and claims data-, predictive models can help physicians and researchers make informed decisions about trial design, patient selection, and potential treatment outcomes.
Predictive analytics is particularly valuable because it can integrate multiple data modalities, such as clinical, biological, genomic, biomarker, and imaging data. The ability to combine and analyze this wealth of information is central to predictive analytics’ potential to revolutionize the clinical trial process.
As the healthcare landscape continues to evolve, predictive analytics will play an increasingly central role in the clinical trial process, driving innovation and improving patient outcomes6.
As with any emerging technology, the adoption of predictive analytics in clinical trials requires collaboration between pharmaceutical companies, regulatory agencies, and healthcare providers. However, the potential benefits are too great to ignore. With predictive analytics at the helm, the future of clinical trials looks promising, offering a path to faster, safer, and more effective drug development.
By leveraging on machine learning, SOPHiA DDM™ facilitates the integration and standardization of diverse data modalities – including but not limited to clinical, biological, radiomics, genomics, and digital pathology data – generating powerful insights to support you in accelerating drug development.
Our multimodal AI data analytics helps you optimize your clinical trial and enhance your post-launch access strategy, by predicting patient response to treatment, disease progression, risk of developing adverse events, and supporting treatment decision-making.
A great example of how SOPHiA GENETICS is spearheading innovation in cancer research by applying predictive analytics is the collaboration with UroCCR, the French Kidney Cancer Research Network, to develop a multimodal machine-learning model for predicting post-operative outcomes for individuals facing renal cell carcinoma (RCC). Using real-world prospective data from the UroCCR network, one of the world’s largest collaborative kidney cancer databases, this study showed that the AI model co-constructed by SOPHiA GENETICS and UroCCR provided a strong prediction for postoperative outcomes, outperforming the predictive performance of most usual risk scores. The results of this study have recently been published in npj Precision Oncology.
With a global network of 780+ institutions, across 70+ countries, and over 1.8 million genomics profiles analyzed to date, the SOPHiA DDM™ Platform accelerates the advances in the field of precision medicine. To learn more about SOPHiA DDM™ for Multimodal and our flagship programs visit our page.
The posters provided real-world data insights on the genomic landscape of specific biomarkers associated with lung and prostate cancers, in a subset of European countries.
The first poster spotlight features Stefano Cheloni, Senior Bioinformatician - Tertiary Analysis at SOPHiA GENETICS, presenting the poster titled: “Real-world insights from France, Italy, Spain, and Austria for the investigation of common (exons 19,21) and rare (exons 18, 20) EGFR mutations in lung cancer”.
The identification of exon-specific EGFR mutations can guide the appropriate treatment of lung cancer with tyrosine kinase inhibitors (TKIs) or alternative targeted therapies. This project aimed to explore the landscape of next-generation sequencing (NGS) testing practices for EGFR mutations in clinical practice in a subset of European countries, to determine the potential number of individuals with lung cancer that could benefit from treatment with TKIs or alternative targeted therapies.
Read the EGFR poster through here.
Watch the spotlight below:
In our second poster spotlight, Adrian Janiszewski, Manager Bioinformatician - Team Lead Tertiary Analysis at SOPHiA GENETICS, showcases a poster titled: “NGS testing practices and molecular profiles of BRCA1/2 in prostate cancer: Real-world insights from France, Italy, Spain, and Austria”.
BRCA1/2 testing can inform which prostate cancer patients might respond to PARP inhibitors (PARPi). Having a better understanding of BRCA1/2 testing practices in the real-world setting can provide valuable insights into identifying gaps and opportunities to improve the identification of metastatic prostate cancer (mPC) patients who may benefit from PARPi treatment. In this study we investigated NGS testing practices results across specific European countries.
Read the BRCA 1 & 2 poster through here.
Watch the spotlight below:
We would like to warmly thank Stefano and Adrian for their insightful presentations and for sharing the key takeaways of these projects.
Learn more about SOPHiA DDM™ for BioPharma, by visiting the page.
DISCLAIMER:
All product and company names are trademarks™ or registered trademarks of their respective holders. Use of them does not imply any affiliation with or endorsement by them. SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
Explore this infographic summary to learn more about the machine learning model developed by Margue et al. for the prediction of disease-free survival in patients undergoing surgery for renal cell carcinoma.

Margue G, et al. UroPredict: Machine learning model on real-world data for prediction of kidney cancer recurrence (UroCCR-120). NPJ Precis Oncol. 2024 Feb 23;8(1):45.
SOPHiA GENETICS multimodal analysis technology and concepts in development. May not be available for sale.
Multimodal healthcare datasets synergistically integrate diverse data modalities such as genomic, clinical, radiomic, proteomic, and biological data, to provide comprehensive insights into human biology and medical conditions. Multimodal datasets have the potential to predict outcomes more accurately and informatively than the sum of their parts (Fig. 1).
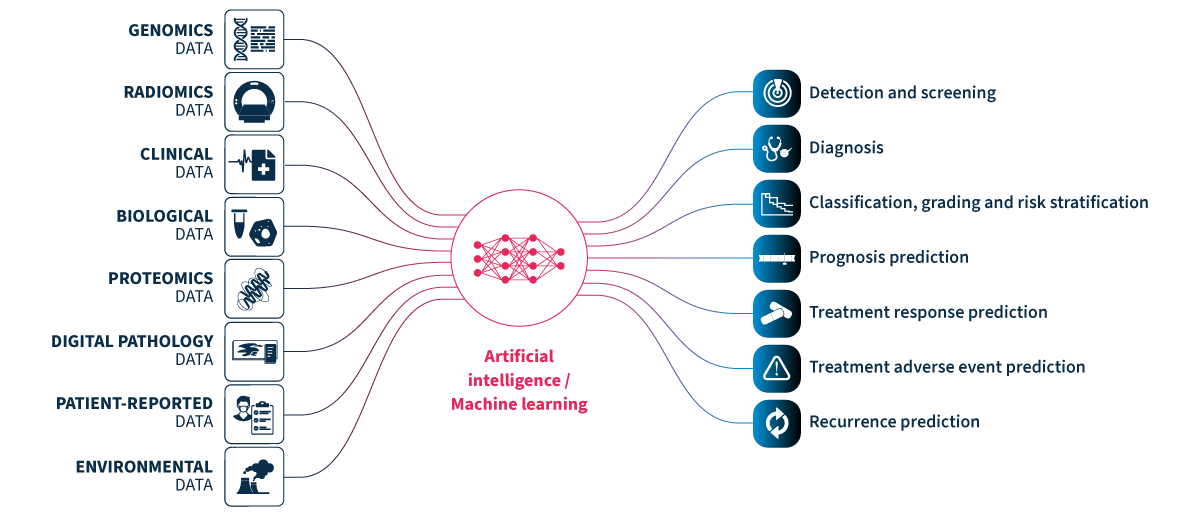
Figure 1.Multimodal healthcare data integrated and analyzed by artificial intelligence (AI)/machine learning can provide useful information for healthcare professionals to use to improve patient care.
Genomics data.
Radiomics data include x-rays, CT scans, MRI scans, ultrasound images, and mammograms.
Clinical and biological data from electronic health records include patient histories, demographics, notes, diagnosis codes, procedure codes, laboratory results, and vital signs.
Proteomics data.
Digital pathology data.
Patient-reported data includes questionnaires and health journals, as well as data from wearable devices monitoring heart rate, sleep patterns, and activity levels, and implantable devices such as pacemakers, insulin pumps, and continuous blood glucose monitors.
Environmental data includes air quality and location data.
New data-driven technologies powered by novel ways of linking and analyzing patient data are set to transform the way that healthcare is delivered.1 Healthcare professionals routinely make use of multiple sources of data to arrive at a diagnosis and to decide on patient management.2 However, a significant level of expertise is required for an in-depth understanding of even a single data type (e.g. radiological images) such that it is unfeasible for individual healthcare professionals to master all areas. AI/machine learning technologies can be leveraged to bring together and analyze multimodal healthcare data, breaking data silos and creating robust and accurate predictive models.3 With the appropriate guidance around decision-making and communication, the valuable insights gained from these predictive models have the potential to support healthcare professionals to improve patient care.
Machine learning technologies can integrate data from disparate multimodal sources to provide a holistic understanding of patients’ health and medical conditions. Data are combined from multiple modalities with the aim of extracting complementary information to power predictive models that can find relationships between different variables/features that are not clearly visible or known by healthcare professionals. Indeed, multimodal data fusion models have consistently shown to provide increased accuracy (1.2-27.7% higher) and performance (AUC 0.02-0.16 higher) than models that utilize data from single modalities for the same task.4
Oncology is one of the medical specialties that most commonly leverages multimodal methods for clinical decision support.5 Machine learning technologies have the potential to explore complex and diverse data to support healthcare professionals from screening to treatment (including relapse).6 Identification of risk factors can support non-invasive patient screening and preventive care.3 Detection of patterns in easily accessible data can help identify diagnostic or prognostic biomarkers to improve patient risk stratification or selection for clinical trials. Identification of predictive signatures of risk factors, adverse treatment reactions, treatment responses, or treatment benefit, can guide decisions around patient management.
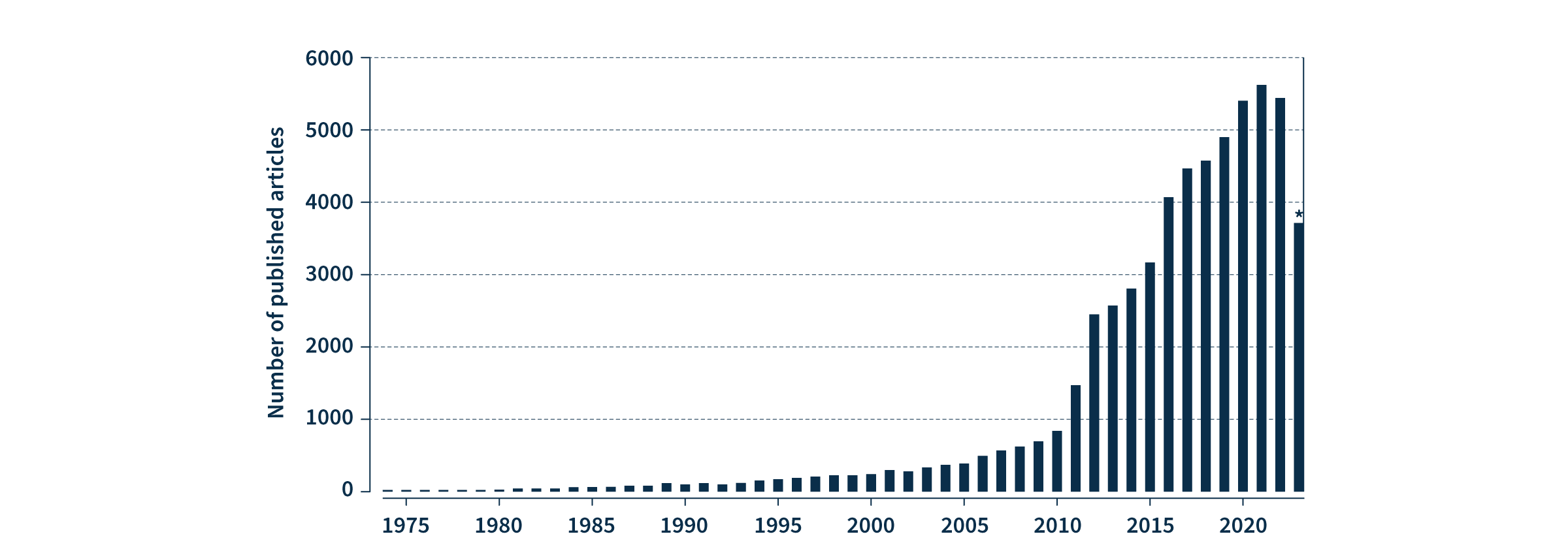
Figure 2. The number of PubMed articles published on multimodal oncology data has dramatically increased in recent years.
PubMed search for ((multimodal) AND (oncology)) OR ((multimodal) AND (cancer)).
*2023 analysis includes data available at time of writing (January-September).
With data privacy and security paramount, multimodal healthcare data can also be leveraged to accelerate advances in medical research, such as the discovery of novel biomarkers and therapeutic targets for drug development, as well as supporting population health management by providing a comprehensive view of health trends and outcomes. The rapid increase in peer-reviewed publications on the topic over the last 13 years demonstrates that the extraordinary value of multimodal oncology data is already recognized by the scientific and medical communities (Fig. 2). Leveraging machine learning to collate and analyze the vast diversity of multimodal data for data-driven precision medicine is on track to drive the next revolution in healthcare.
SOPHiA DDM™ multimodal healthcare analytics will have the potential to break data silos by streamlining the integration of longitudinal oncology data from multiple sources and modalities – including but not limited to genomic, radiomic, digital pathology, biological, and clinical data. The SOPHiA DDM™ Platform uses machine learning-powered analytics to assemble, standardize, and transform multimodal data into accessible data-driven insights, facilitating the identification of multimodal predictive signatures, as well as treatment response patterns and trends. To learn more and get in touch, visit the webpage.
Product in development – Technology and concepts in development. May not be available for sale.
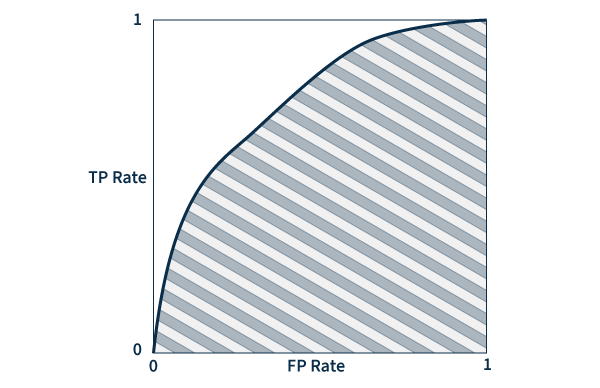
Area under the ROC curve (AUC) – A ROC (receiver operating characteristic) curve is a graph that plots true and false positive rates to demonstrate the performance of a model. AUC measures the area underneath the ROC curve to provide an aggregate measure of performance. AUC values range between 0 and 1, with a score of 0 meaning that all predictions are wrong, and a score of 1 meaning that all predictions are 100% correct. Essentially, AUC represents the probability that a positive result is truly positive and a negative result is truly negative.
Omics data – Large-scale information related to the biology of organisms.
Digital pathology images – Scanned images of tissue samples on glass slides.
Companion diagnostics (CDx) are medical devices, specifically an in vitro diagnostic device (IVD), providing important information regarding the safe and effective use of therapeutics. The Food and Drug Administration (FDA) ascribes them three crucial functions: 1) identify patients more likely to benefit from a therapeutic; 2) determine patients at increased risk of serious side effects; and 3) monitor treatment responses for the purpose of adjusting dosage or regimens to improve safety and effectiveness. In our estimation, CDx act as a compass that directs the healthcare provider to the most appropriate treatment for each patient1.
The inception of CDx can be traced back to 1998, when the FDA granted concurrent approval for trastuzumab, a targeted cancer drug, and HercepTest™, a HER2 immunohistochemical assay. This milestone marked the birth of the drug-diagnostic co-development model, a transformative approach that has since witnessed consistent and substantial adoption2.
However, over the next 14 years, CDx advancement was slow, with the majority of new approvals occurring only in the past decade. In fact, from 1998 to 2012, approximately 20 new CDx were approved, whereas from 2013 to 2023, that number rose to 1343.
Today, approximately 50% of all new molecular entity (NME) approvals in oncology have an associated CDx or biomarker listed in the label required for safe and effective use (based on FDA approvals from 2021 and 2022 of NME in oncology3). Despite the historical tendency toward oncology products, applications in rare diseases and metabolic syndromes are evolving, paving the way for CDx to become an intrinsic part of precision medicine clinical trials across many indications.
The use of a companion diagnostics strategy in clinical trials, which we define here as using one or more biomarkers to pre-select and enroll patients more likely to respond to the experimental therapy, is commonly employed in oncology. In these trials, identification and pre-selection have significant advantages, allowing smaller patient groups to power the statistical analysis, potentially reducing overall costs, and increasing the likelihood of approval4.
But while a CDx strategy makes regulatory approval of a cancer drug more likely, it can simultaneously add complexity to the process:
Despite the evident complexity, the widespread adoption of next-generation sequencing (NGS) has made using companion diagnostics and deploying biomarker-driven strategies in clinical trials easier by permiting screening for multiple biomarkers simultaneously. Rather than rely on a one-biomarker-one-test model, NGS permits patients to be screened for eligibility for multiple therapeutics or clinical trials.. Moreover, targeted gene panels and broader approaches, such as comprehensive genomic profiling (CGP) and whole exome sequencing (WES) have further streamlined the development of multi-biomarker-driven CDx.
Returning to our analogy, while CDx acts as the compass, NGS technologies are the roadmap to determine the most suitable treatment for each patient.
There are two well-defined pathways to approval for companion diagnostics:
HercepTest™ followed the first pathway where the physical kit, produced by an IVD manufacturer, received a Pre-market Approval (PMA). In contrast, the precedent for the single-site model was only established much later with the publication of clinical evidence supporting the hypothesis that BRCA-mutated patients were more likely to benefit from treatment with olaparib—a PARP inhibitor first approved for the treatment of advanced ovarian cancer5,6.
Requiring a complex workflow and expert oversight, BRCA analysis was more conducive to the simplicity of the single-site model since the very nature of this pathway streamlines validation – validating a workflow within a single lab is less time-consuming than across multiple labs. This new approach played a pivotal role in rapidly advancing the commercial adoption of NGS applications.
Today, most NGS-powered CDx assays follow the single-site pathway3, creating a new set of challenges. Despite the increased simplicity, this model confines assays to single locations, limiting the capacity for sample analysis, significantly increasing turnaround times, and reducing patient access. In the new age of precision medicine, these limitations are being addressed through a decentralized testing and analysis model supported by technology-agnostic and easy-to-implement workflows.
While direct co-development of a CDx and therapeutic stands as the preferred regulatory model by the FDA, alternative approaches may be utilized due to the inherent challenges of aligning IVD and drug development.
While the development of a CDx and therapeutic are tightly entwined, drug developers and IVD manufacturers remain separate entities with a few exceptions. This requires the carefully selection of partner(s) within the IVD and CDx ecosystem to ensure successful programs.
Many questions must be addressed early in the process, as even suboptimal approaches can significantly delay and impact commercial uptake:
The advent of NGS technologies has heralded a healthcare revolution, propelling us toward data-driven precision medicine. Yet, as we push ahead in developing biomarker-driven applications for a plethora of indications, we face the potential for increased implementation challenges, threatening to complicate the patient journey.
The adoption of a decentralized, globally accessible, intuitive, and technology-agnostic SOPHiA DDM™ Platform, leveraging proprietary algorithms and a vast portfolio of robust NGS-based applications, is well positioned to streamline CDx co-development and implementation, enhancing access to analytically robust solutions without overtaxing healthcare resources.
Our holistic approach to co-development is poised to chart a course toward a more integrated future, arming developers with the necessary data and insights to tackle potential hurdles and maximize the time and resources allocated to clinical research programs.
At SOPHiA GENETICS, our unwavering commitment is to collaboratively engineer deployable solutions that elevate implementation and accessibility in precision testing while streamlining the process of analysis and interpretation. Explore the possibilities of SOPHiA DDM™ for BioPharma by visiting our dedicated page.
References
1 Food and Drug Administration (FDA). In Vitro Companion Diagnostic Devices: Guidance for Industry and Food and Drug Administration Staff. Issued on: August, 2014. Accessed on: October, 2023. Retrieved from: https://www.fda.gov/media/81309/download
2 Jørgensen JT. Drug-diagnostics co-development in oncology. Front Oncol. 2014;4:208. doi: 10.3389/fonc.2014.00208
3 FDA. List of Cleared or Approved Companion Diagnostic Devices (In Vitro and Imaging Tools). Accessed on: October 2023. Retrieved from: https://www.fda.gov/medical-devices/in-vitro-diagnostics/list-cleared-or-approved-companion-diagnostic-devices-in-vitro-and-imaging-tools
4 Biotechnology Industry Organization (BIO). Clinical Development Success Rates and Contributing Factors 2011–2020. Accessed on: October 2023. Retrieved from: https://go.bio.org/rs/490-EHZ-999/images/ClinicalDevelopmentSuccessRates2011_2020.pdf
5 Ledermann J, et al. Olaparib maintenance therapy in patients with platinum-sensitive relapsed serous ovarian cancer: a preplanned retrospective analysis of outcomes by BRCA status in a randomised phase 2 trial. Lancet Oncol. 2014;15(8):852-61. doi: 10.1016/S1470-2045(14)70228-1.
6 Deeks ED. Olaparib: first global approval. Drugs. 2015;75(2):231-240. doi: 10.1007/s40265-015-0345-6
7 Hanna TP, et al. Mortality due to cancer treatment delay: systematic review and meta-analysis. BMJ. 2020 Nov 4;371:m4087. doi: 10.1136/bmj.m4087
Precision medicine, also known as personalized medicine, aims to enhance healthcare quality by tailoring treatments to each person's unique genetic makeup, environment, and lifestyle. While a fully individualized approach to medicine is still a work in progress, the recognition that patient heterogeneity influences treatment effectiveness is not new1.
Historically, medicine has heavily relied on trial-and-error strategies for discovering, developing, and testing new treatments targeted at specific indications. This disease-centered approach resulted in predetermined standard therapies tailored to the “average patient.” While this one-size-fits-all approach has succeeded in many indications, it also carries significant drawbacks, particularly when dealing with complex diseases such as cancer or inherited disorders. In these cases, the risk of adverse side effects (e.g., toxicity) and reduced therapeutic response often result in poorer patient prognoses and quality of life1.
Precision medicine represents a patient-centric paradigm shift, acknowledging each individual's uniqueness while using real-world data and advanced statistics to guide the discovery-to-treatment process. For instance, pharmacogenomics requires us to look at each patient genomic data individually and in the context of others, enabling stratification into cohorts for predicting treatment responses2.
Success in precision medicine hinges on the ability to derive meaningful insights from large patient datasets. Fostering data diversity has the potential to further advance progress in this area1,3.
Recent technological advances have made precision medicine more accessible and impactful than ever before. Next-generation sequencing (NGS) has become more affordable, transforming it from a research-focused technology into a tangible clinical reality. This progress was further propelled by the widespread adoption of electronic health records (EHRs) and laboratory information management systems (LIMS), which not only facilitate population-scale research but also enable the use of clinical decision support tools for the delivery of targeted therapies to individual patients4.
The ability to identify genetic biomarkers and assess variant pathogenicity has grown significantly in the past decade. This has not only revolutionized patient diagnosis but also transformed drug development. A pivotal moment was the approval of imatinib by the FDA in 2001, the first small molecule targeted therapy for chronic myeloid leukemia (CML)5. By inhibiting the BCR-ABL fusion protein, imatinib was shown to effectively slow the progression of CML from chronic phase to blast crisis, making it the first of its kind.
This groundbreaking milestone paved the way for the approval of many other targeted therapies, such as gefitinib targeting EGFR alterations associated with NSCLC (2003) and trastuzumab for HER2-positive breast cancer (2006). The pace of new targeted drug approvals continues to accelerate year after year, heralding a promising era of precision medicine6.
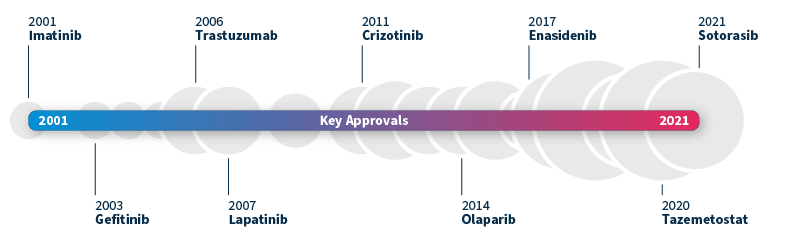
Timeline of FDA-approved targeted therapies in cancer. Grey bubbles represent the relative number of approvals per year. Data source: Waarts et al 2022.
To achieve a truly personalized approach to medicine, the harmonization of translational and precision medicine is paramount. This coordination between early mechanism-based drug development and late-stage patient-centric approaches gives rise to an end-to-end biomarker-guided process, allowing us to optimize treatment strategies for patient cohorts right from the outset7.
Known as translational precision medicine, this emerging concept brings a fresh perspective to the translational gap, calling for a broader scope beyond a purely genetic-based definition of biomarkers and introducing a multimodal approach by taking into account a wider range of healthcare variables. To make this new concept a reality, significant technological progress is required in several key areas8:
By addressing these critical areas of advancement, we can pave the way for a future where each patient receives personalized treatments tailored to her or his unique needs and characteristics. The pursuit of translational precision medicine promises to revolutionize healthcare, offering improved patient outcomes and transforming the landscape of medical research and development.
Powered by proprietary algorithms and enriched with data from 750+ institutions*, the SOPHiA DDM™ Platform accelerates advances in the field of precision medicine. Its core mission centers on empowering clinical researchers across healthcare and biopharma spheres alike.
To learn more about SOPHiA DDM™ BioPharma Solutions for biomarker-centric discovery, development, and application deployment, visit our page.
* The number of institutions represents active customers who have generated revenue through the SOPHiA DDM™ Platform usage or Alamut™ Visual Plus licenses as of September 30, 2022.
SOPHiA GENETICS products are for Research Use Only and not for use in diagnostic procedures unless specified otherwise.
SOPHiA DDM™ Dx Hereditary Cancer Solution, SOPHiA DDM™ Dx RNAtarget Oncology Solution and SOPHiA DDM™ Dx Homologous Recombination Deficiency Solution are available as CE-IVD products for In Vitro Diagnostic Use in the European Economic Area (EEA), the United Kingdom and Switzerland. SOPHiA DDM™ Dx Myeloid Solution and SOPHiA DDM™ Dx Solid Tumor Solution are available as CE-IVD products for In Vitro Diagnostic Use in the EEA, the United Kingdom, Switzerland, and Israel. Information about products that may or may not be available in different countries and if applicable, may or may not have received approval or market clearance by a governmental regulatory body for different indications for use. Please contact us to obtain the appropriate product information for your country of residence.
All third-party trademarks listed by SOPHiA GENETICS remain the property of their respective owners. Unless specifically identified as such, SOPHiA GENETICS’ use of third-party trademarks does not indicate any relationship, sponsorship, or endorsement between SOPHiA GENETICS and the owners of these trademarks. Any references by SOPHiA GENETICS to third-party trademarks is to identify the corresponding third-party goods and/or services and shall be considered nominative fair use under the trademark law.